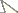(C) 2011 Priscilla Cardim Scacchetti. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
For reference, use of the paginated PDF or printed version of this article is recommended.
Conventional (Giemsa, C-Banding, Ag-NORs, CMA3) and molecular (5S rDNA, 18S rDNA, telomeric sequences) cytogenetic studies were carried out in specimens of ten distinct fish populations of the genus Gymnotus (Gymnotus sylvius Albert and Fernandes-Matioli, 1999, Gymnotus inaequilabiatus Valenciennes, 1839, Gymnotus pantherinus Steindachner, 1908, and G. cf. carapo Linnaeus, 1758) from different Brazilian hydrographic basins. Gymnotus sylvius presented a diploid number of 40 chromosomes (22m+12sm+6st), Gymnotus pantherinus presented 52 chromosomes (32m+18sm+2st), while Gymnotus inaequilabiatus (42m+10sm+2a)and Gymnotus cf. carapo (38m+12sm+4st) presented 54 chromosomes. The C-banding technique revealed centromeric marks in all chromosomes of all species. Besides that, conspicuous blocks of heterochromatin were found interstitially on the chromosomes of Gymnotus inaequilabiatus, Gymnotus cf. carapo, and Gymnotus pantherinus. All four species showed single nucleolus organizing regions confirmed by results obtained through Ag-NORs and FISH experiments using 18S rDNA probes, which showed the NORs localized on the first chromosome pair in Gymnotus inaequilabiatus, Gymnotus cf. carapo, and Gymnotus pantherinus, and on pair 2 in Gymnotus sylvius. CMA3 staining revealed additional unrelated NORs marks in Gymnotus sylvius and Gymnotus pantherinus. The 5S rDNA probes revealed signals on one pair in Gymnotus sylvius and two pairs in Gymnotus pantherinus; Gymnotus inaequilabiatus had about seventeen pairs marked, and Gymnotus cf. carapo had about fifteen pairs marked. It is considered that the high amount of heterochromatin identified in the chromosomes of Gymnotus inaequilabiatus and Gymnotus cf. carapo could have facilitated the dispersion of 5S rDNA in these species. Interstitial signals were detected on the first metacentric pair of Gymnotus sylvius by telomeric probes (TTAGGG)n indicating the possible occurrence of chromosomal fusions in this species. The present study reveals valuable cytotaxonomic markers for this group and allows a more precise evaluation of the processes involved in the karyotype differentiation and the interrelationships among different species of the genus Gymnotus.
FISH, rDNA, cytogenetics, heterochromatin, chromosomal rearrangements
Fish species belonging to the order Gymnotiformes,
usually known as “tuviras”, “electric fish”, or “banded knife-fishes”,
constitute a group endemic to the Neotropical region (
Gymnotidae is currently composed of the genus Gymnotus, with 35 valid species, and Electrophorus Gill, 1864 with only one valid species (
The available cytogenetics data for Gymnotus species evidence a high karyotypic diversity characterized by different diploid numbers observed in some species, as in Gymnotus carapo Linnaeus, 1758 and Gymnotus inaequilabiatus Valenciennes, 1839with 54 chromosomes; Gymnotus sylvius Albert and Fernandes-Matioli, 1999 which shows 40 chromosomes; Gymnotus pantherinus Steindachner, 1908 with 52 chromosomes, and Gymnotus capanema
The current study was carried out aiming to broaden the cytogenetic data available for Gymnotus, mapping the distribution of ribosomal sites and telomeric DNA sequences on the chromosomes of different species of this genus. The data obtained will allow a better understanding of the mechanisms involved in the process of karyotypic differentiation and diversification of this fish group.
Material and methodsFour fish species of Gymnotus sampled throughout the different components of the Brazilian hydrographic river basins were cytogenetically analyzed (Figure 1 and Table 1). After analysis, the specimens were deposited in the fish collection of the Laboratório de Biologia e Genética de Peixes (LBP), Universidade Estadual Paulista, at Botucatu, São Paulo, Brazil.
Map of Brazil showing the collection sites of species and populations of Gymnotus analyzed. 1 Miranda River, Passo do Lontra – MT, Gymnotus cf. carapo 2 Campo Novo River, Bauru – SP, Gymnotus sylvius and Gymnotus inaequilabiatus 3 Água da Madalena River, Botucatu – SP, Gymnotus sylvius and Gymnotus inaequilabiatus 4 Araquá River, Botucatu – SP, Gymnotus sylvius and Gymnotus inaequilabiatus 5 Mogi-Guaçu River, Pirassununga – SP, Gymnotus sylvius and Gymnotus inaequilabiatus; 6. Aguapeú River, Mongaguá – SP, Gymnotus pantherinus.
Specimens of Gymnotus analyzed. LBP – deposit voucher number at the fish collection of the Laboratório de Biologia e Genética de Peixes, Instituto de Biociências de Botucatu, UNESP. F – females, M – males.
| Species | LBP | Sample Localities | F | M | Coordinates |
|---|---|---|---|---|---|
| Gymnotus sylvius | 11160 | Água da Madalena - Botucatu-SP River | 20 | 14 | S22°59.25', W48°25.40' |
| Gymnotus sylvius | 11155 | Araquá – Botucatu-SP River | 02 | - | S22°47.13', W48°28.89' |
| Gymnotus sylvius | 11163 | Campo Novo- Bauru-SP River | 01 | 01 | S22°23.07', W49°00.55' |
| Gymnotus sylvius | 11161 | Mogi-Guaçu - Pirassununga-SP River | - | 01 | S21°55.50', W47°22.29' |
| Gymnotus inaequilabiatus | 11154 | Água da Madalena - Botucatu-SP River | 02 | 07 | S22°59.25', W48°25.40' |
| Gymnotus inaequilabiatus | 11158 | Araquá – Botucatu-SP River | 04 | 02 | S22°47.13', W48°28.89' |
| Gymnotus inaequilabiatus | 11152 | Campo Novo - Bauru-SP River | 06 | 13 | S22°23.07', W49°00.55' |
| Gymnotus inaequilabiatus | 11156 | Mogi-Guaçu - Pirassununga-SP River | 06 | 17 | S21°55.50', W47°22.29' |
| Gymnotus pantherinus | 11153 | Aguapeú - Mongaguá-SP River | 03 | 02 | S24°06.40', W46°43.00' |
| Gymnotus cf. carapo | 9836 | Miranda - Pantanal-MSRiver | 03 | 02 | S19°34.34', W57°02.17' |
The fishes were euthanized with a lethal dose of
benzocaine before the procedures of chromosome preparation. Mitotic
chromosome preparations were carried out according to
Molecular cytogenetic analysis involved the use of GC-specific fluorochrome Chromomycin A3 (CMA3) (
Cytogenetic analysis performed in representatives of four Gymnotus fish species evidenced an expressive variation in the diploid number among the species, despite the conservative karyotypic feature among the representatives of the populations. Gymnotus sylvius presented 40 chromosomes (Fig. 2a); Gymnotus inaequilabiatus and Gymnotus cf. carapo presented 54 chromosomes (Figs 3a, 5a), and Gymnotus pantherinus presented 52 chromosomes (Fig. 4a). Data are summarized in Table 2.
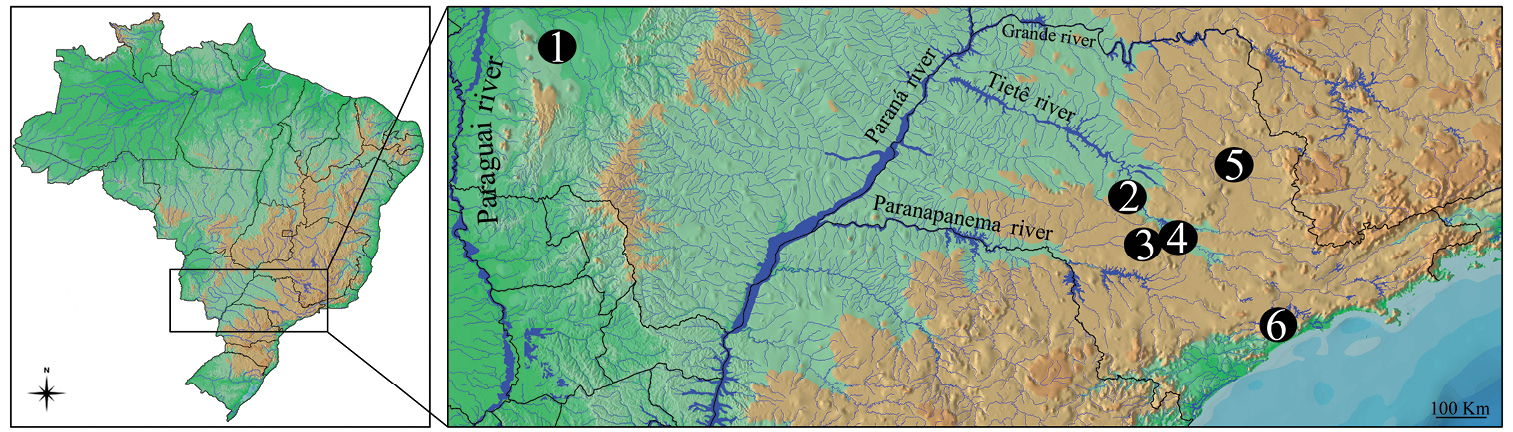
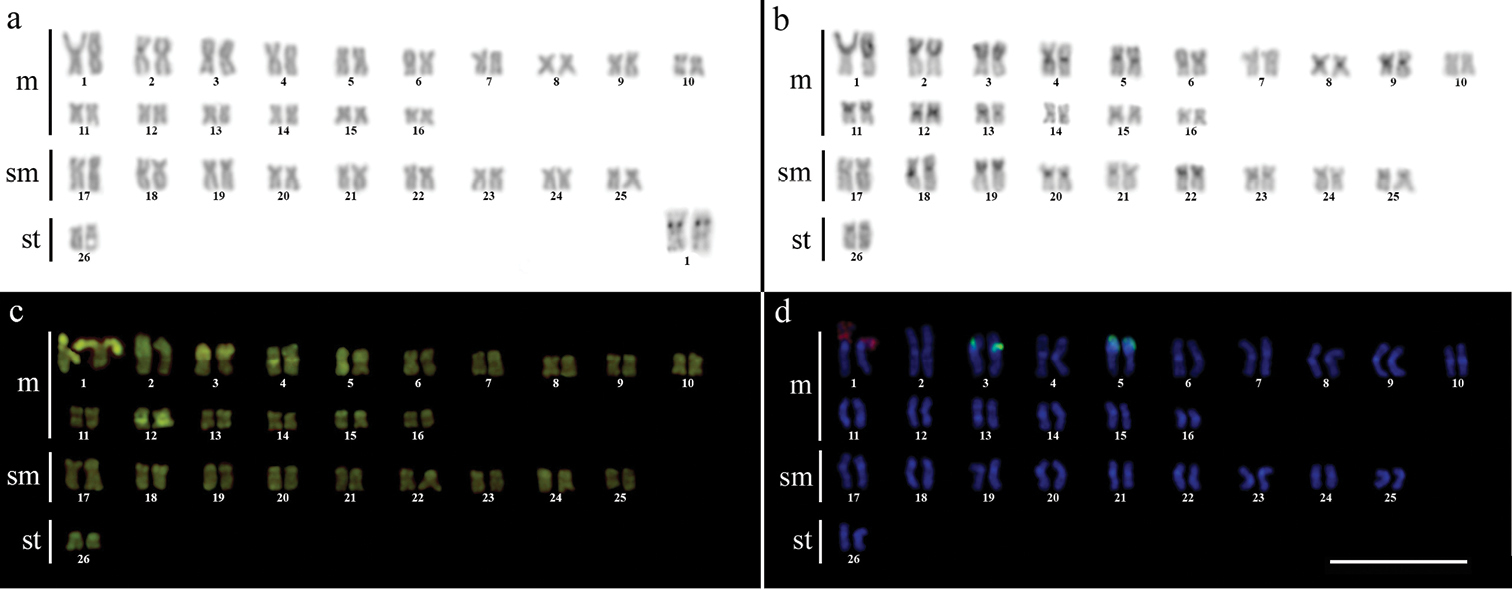
Karyotype of Gymnotus sylvius after (a) conventional Giemsa staining, (b) C-banding, (c) CMA3 fluorochrome staining, (d) double FISH with 5S rDNA (red) and 18S rDNA (green) probes. Bar = 10µm.
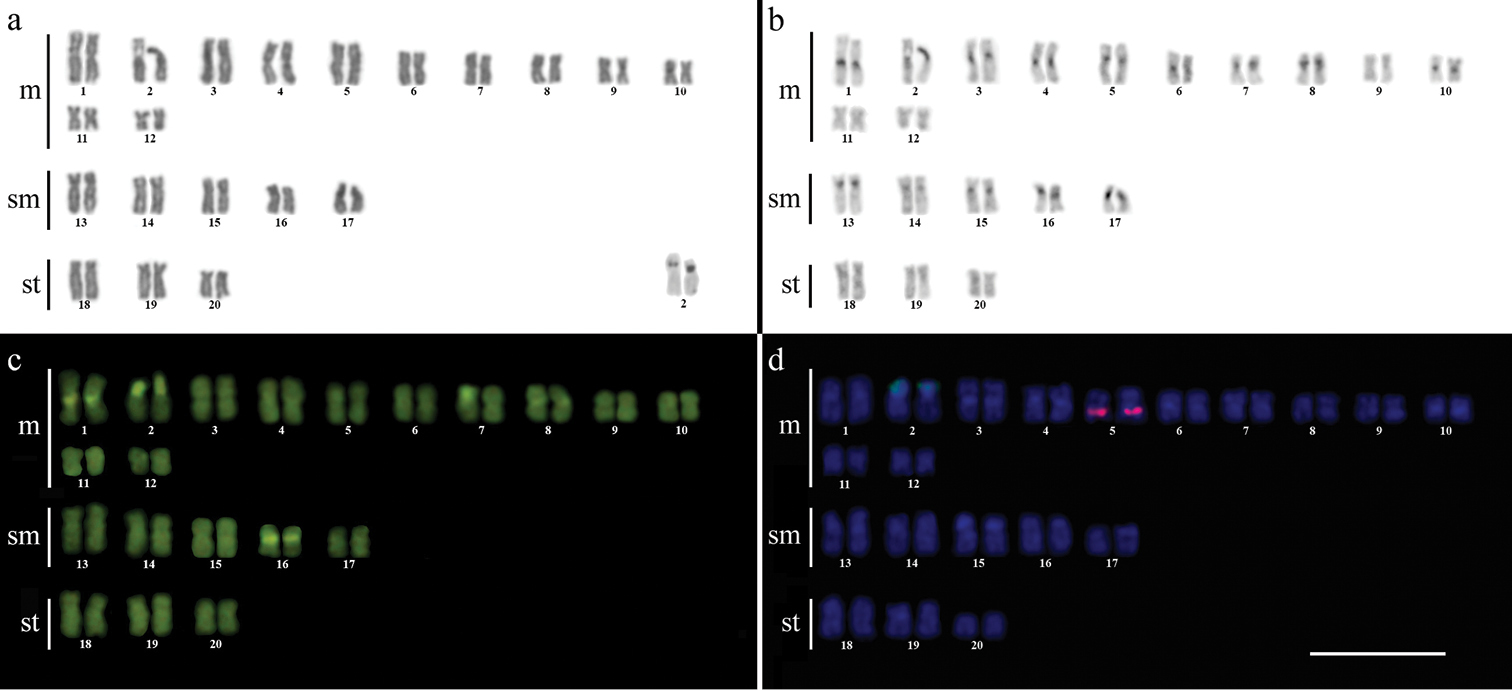
Karyotype of Gymnotus inaequilabiatus after (a) conventional Giemsa staining, (b) C-banding, (c) CMA3 fluorochrome staining, (d) double FISH with 5S rDNA (red) and 18S rDNA (green) probes. Bar = 10µm.
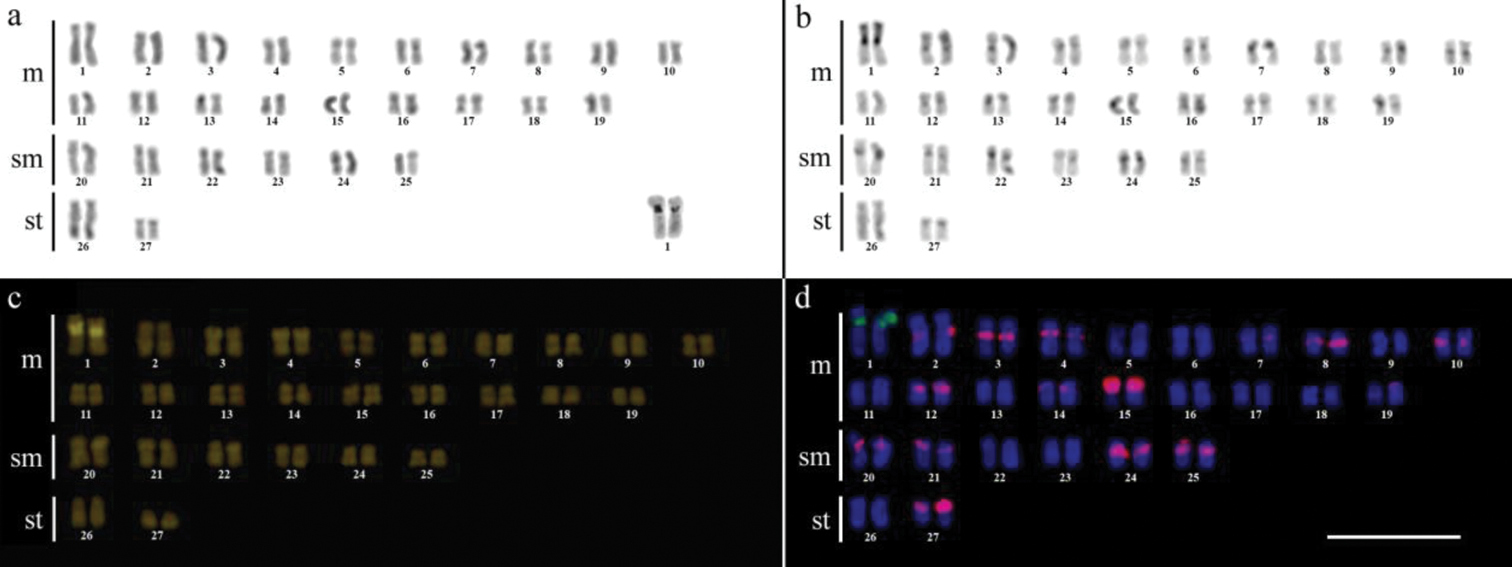
Karyotype of Gymnotus pantherinus after (a) conventional Giemsa staining, (b) C-banding, (c) CMA3 fluorochrome staining, (d) double FISH with 5S rDNA (green) and 18S rDNA (red) probes. Bar = 10µm.
Karyotype of Gymnotus cf. carapo after (a) conventional Giemsa staining, (b) C-banding, (c) CMA3 fluorochrome staining, (d) double FISH with 5S rDNA (red) and 18S rDNA (green) probes. Bar = 10µm.
Cytogenetic data on four species of Gymnotus. ITS – Interstitial Telomeric Sites; (I) – Interstitial mark.
| Species | 5S rDNA | 18S rDNA | ITS | CMA3 | Karyotypic formulae |
|---|---|---|---|---|---|
| Gymnotus sylvius | Pair 4 | 2 (I) | Pair 1 | Pairs 1, 2 and 16 | 22m+12sm+6st |
| Gymnotus inaequilabiatus | Up to17 pairs | 1 (I) | -- | Pair 1 | 42m+10sm+2a |
| Gymnotus pantherinus | Pairs 3 and 5 | 1 (I) | -- | Pairs 1, 3, 4 and 12 | 32m+18sm+2st |
| Gymnotus cf. carapo | Up to 15 pairs | 1 (I) | -- | Pair 1 | 38m+12sm+4st |
The C-banding technique revealed significant differences in the distribution patterns of heterochromatin among the analyzed species. All populations of Gymnotus sylvius showed small amounts of constitutive heterochromatin restricted to the centromeric areas of all chromosomes, and also blocks coincident with NORs (Fig. 2b). In Gymnotus inaequilabiatus (Fig. 3b), Gymnotus pantherinus (Fig. 4b) and Gymnotus cf. carapo (Fig. 5b), besides centromeric and pericentromeric marks, it was possible to observe conspicuous interstitial blocks of heterochromatin in some chromosomes. No numerical or structural polymorphisms related to the presence of supernumerary or sex chromosomes were detected in the samples analyzed.
The impregnation by silver nitrate evidenced that all species and populations of Gymnotus analyzed hold a simple pair of chromosomes bearing NORs. The populations of Gymnotus sylvius showed signalsat the interstitial region on the short arms of chromosome pair 2 (highlighted in Fig. 2a). The representatives of the other species showed their ribosomal sites located in the interstitial position on the short arms of chromosome pair number 1 (highlighted in Figs 3a–5a). The use of 18S rDNA probe confirmed the results achieved with silver nitrate staining (Figs 2d–5d), while the hybridization with 5S rDNA probes localized this gene in the pericentromeric position of pair number 4 in the representatives of Gymnotus sylvius populations (Fig. 2d); in two chromosome pairs (numbers 3 and 5) in Gymnotus pantherinus (Fig. 4d); in up to 17 chromosomal pairs in the representatives of Gymnotus inaequilabiatus, and in up to 15 pairs in Gymnotus cf. carapo (Figs 3d, 5d). The coloration with fluorochrome CMA3 in Gymnotus inaequilabiatus and Gymnotus cf. carapo marked only the pair bearing the NORs (Figs 3c, 5c), while Gymnotus sylvius and Gymnotus pantherinus showed additional marked pairs besides the chromosomes bearing NORs (Figs 2c, 4c).
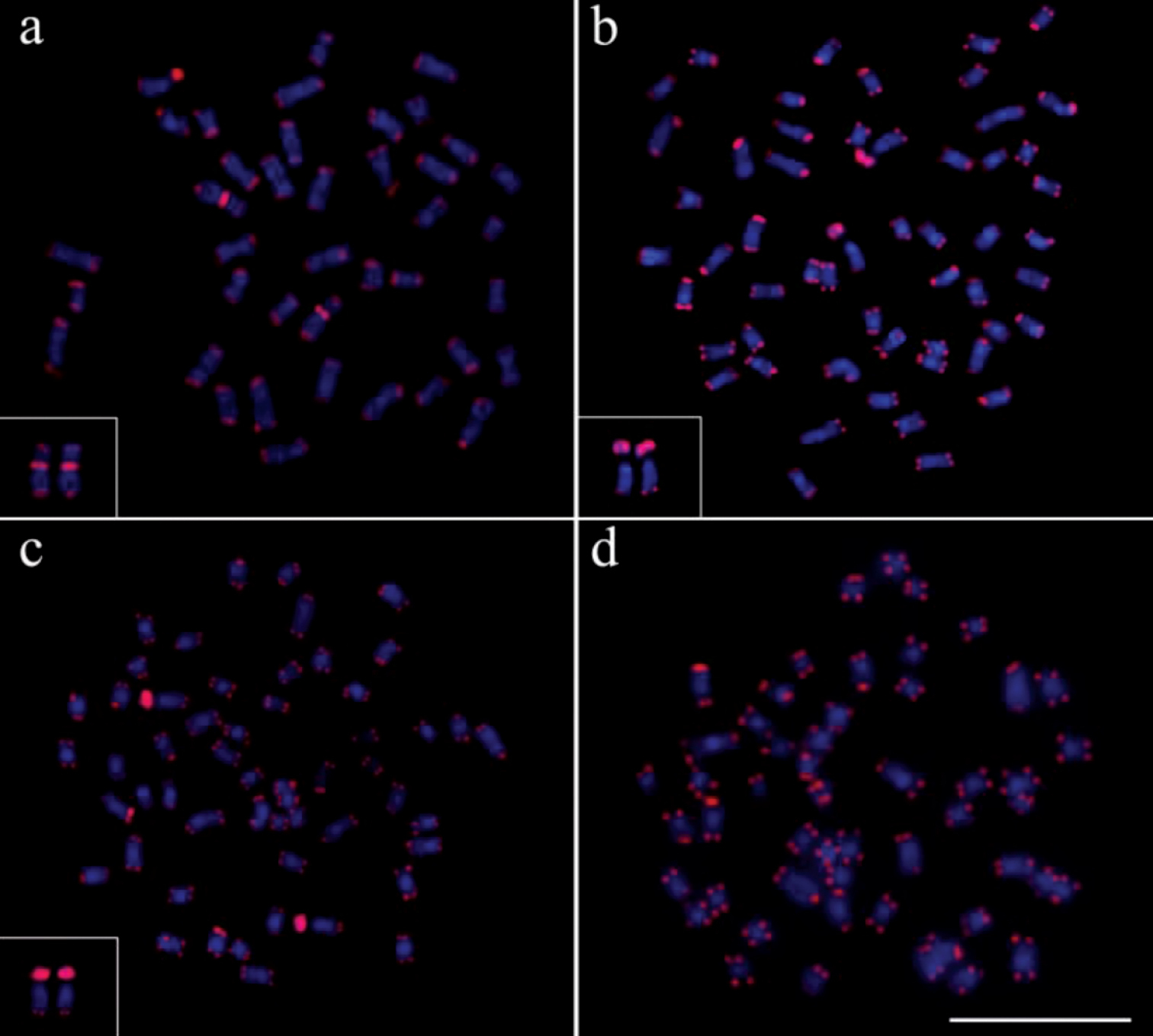
The use of telomeric probes (TTAGGG)n evidenced signals in the terminal position of all chromosomes in all populations analyzed (Fig. 6). Additionally, conspicuous marks were found along the nucleolar regions in the specimens of Gymnotus inaequilabiatus (Fig. 6b) and Gymnotus cf. carapo (Fig. 6c). Besides that, interstitial telomeric sites (ITS) were observed in the chromosomes of pair number 1 in Gymnotus sylvius (Fig. 6a). All data are summarized in Table 2 and represented in an ideogram in Figure 7.
Distribution pattern of telomeric sites in metaphases of the representatives of the four species of Gymnotus analyzed. (a) Gymnotus sylvius (featured interstitial telomeric sites – ITS), (b) Gymnotus inaequilabiatus, (c) Gymnotus cf. carapo and (d) Gymnotus pantherinus. Bar = 10µm.
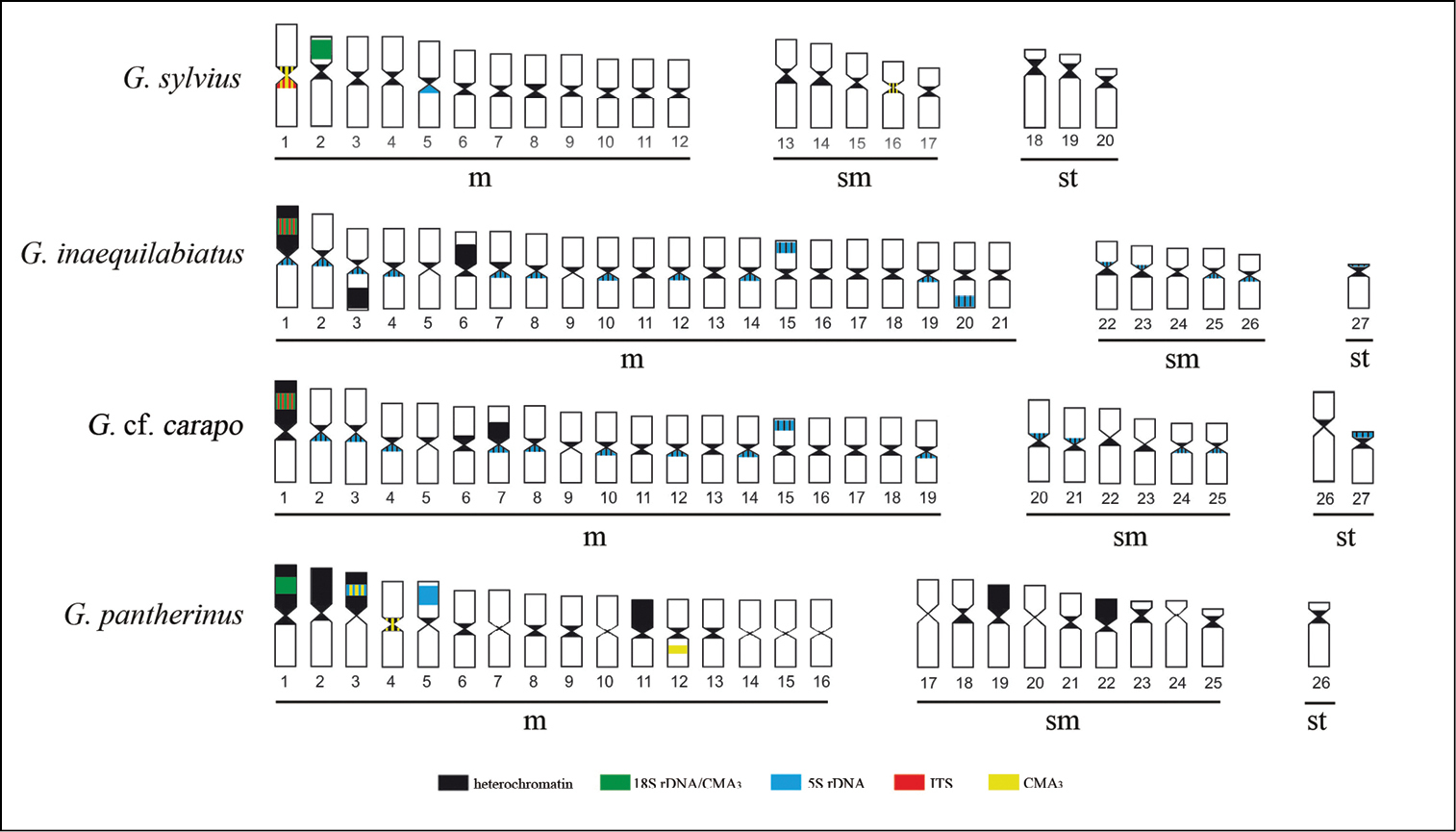
Ideogram showing the hybridization patterns described in this paper. The overlapping signals are represented simultaneously by the respective colors (a) Gymnotus sylvius, (b) Gymnotus inaequilabiatus, (c) Gymnotus cf. carapo and (d) Gymnotus pantherinus.
Available cytogenetic data on the genus Gymnotus
evidence the occurrence of high karyotypic diversity among the species,
notably related to diploid number and karyotypic formulae, ranging
from 34 chromosomes in Gymnotus capanema up to 54 chromosomes in Gymnotus inaequilabiatus (
The karyotype diversity found in the species may be related to the fact that the representatives of Gymnotus
are generally endemic organisms living in headwaters, which do not
migrate long distances. Such characteristic may act to reduce gene flow
among different populations, even in the same hydrographic basin,
resulting in differences between populations of the same species, as
found in samples of Characidium Reinhardt, 1867 (
The karyotypic identity observed among populations inside the species of Gymnotus
reinforces the postulate that cytogenetic characteristics could be
considered an important tool for taxonomic diagnostic of species in this
fish group (
The differences related to the number and morphology of
the chromosomes found among the species suggest the occurrence of
structural and numerical rearrangements during the process of
differentiation.
In the current study, the probes used for telomeric
sequence (TTAGGG)n revealed signals of hybridization on the extremities
of all chromosomes in all populations analyzed (Figure 6). However, interstitial telomeric sites (ITSs) were observed in the chromosomes of Gymnotus sylvius.
The presence of these ITSs in some chromosomes could be an indicative
of recent centric fusion events, as previously discussed by
The occurrence of ITS in some chromosomes of Gymnotus sylvius, as well as its absence in Gymnotus pantherinus,
could be attributed to different factors, such as the occurrence of
differences in the type of chromosomal rearrangements, the plasticity
of the telomeric sequences or the divergence time of the species, which
originated modifications in the sequences and possibly made them
undetectable by the FISH technique. In a phylogenetic reorganization of
the Gymnotiformes based on molecular and cytogenetic data,
The identification of nucleolus organizer regions in the
four species analyzed through silver nitrate staining and 18S rDNA
probes revealed only one chromosome pair containing nucleolar sites and
characterizing a simple NORs system, as previously cited (
The use of CMA3 in metaphase chromosomes revealed additional marks to those identified in the ribosomal sites in Gymnotus sylvius and Gymnotus pantherinus, indicating the presence of additional GC-rich sequences.
The distribution patterns of 5S rDNA sequences showed a
peculiar dispersion of these repetitive sites in the karyotypes of the
species in the genus Gymnotus. Considering the great blocks of heterochromatin and the variation in the distribution pattern of 5S rDNA sites in Gymnotus inaequilabiatus, Gymnotus cf. carapo, and even in Gymnotus pantherinus,
it can be considered that this situation might have favored the
occurrence of structural rearrangements in the karyotype of these
species, since heterochromatic areas are more propitious to breaks,
and thus may facilitate the dispersion of this gene sequence. The
absence of large heterochromatic blocks and the presence of 5S rDNA in a
unique pair of chromosomes in Gymnotus sylvius could reinforce this hypothesis.
Recent studies carried out by
Despite the marked karyotypic conservation found inside the populations of Gymnotus species, the results achieved in the current work revealed great differences in the chromosome structure in the species of this genus, indicating that all possible evolution ways passed through the differentiation process of chromosomes.
This research was supported by CNPq, CAPES and FAPESP.