(C) 2011 Valentina Monti. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
For reference, use of the paginated PDF or printed version of this article is recommended.
Cytogenetic and molecular investigations on the holocentric chromosomes of the aphid Macrosiphum euphorbiae (Thomas, 1878)have been carried out using silver staining and C-banding (followed by chromomycin A3 and DAPI staining) in order to improve our knowledge about the structure of aphid chromosomes. The C-banding pattern is peculiar since only the two X chromosomes and a single pair of autosomes presented heterochromatic bands. Silver staining and FISH with the 28S rDNA probe localized the rDNA genes on one telomere of each X chromosome that were also brightly fluorescent after chromomycin A3 staining of C-banded chromosomes, whereas all other heterochromatic bands were DAPI positive. Interestingly, a remarkable nucleolar organizing region (NOR) heteromorphism was present making the two X chromosomes easily distinguishable. Southern blotting and FISH assessed the presence of the (TTAGG)n repeat at the ends of all the Macrosiphum euphorbiae chromosomes. Karyotype analysis showed that all males possessed the X chromosome with the larger amount of rDNA suggesting a non-Mendelian inheritance of the two X chromosomes.
aphid, holocentric chromosomes, telomeres, heterochromatin, NOR heteromorphism
In the last decades classic and molecular cytogenetics
provided an integrated approach for the structural, functional and
evolutionary analysis of aphid holocentric chromosomes (
Interest in aphid cytogenetics is mostly due to the
holocentric/holokinetic structure of their chromosomes that present a
diffused centromeric activity (
Aphids, in view of the ease with which mitotic
chromosome can be obtained from embryonic tissues, represent an ideal
model to better understand the architecture of holocentric chromosomes,
and to work out the differences/similarities with monocentric ones (
Aphid X chromosomes have been studied with great attention since they present several structural constraints (
In order to better understand X chromosome evolution in
aphids, we decided to carry out a cytogenetic analysis of the
holocentric chromosomes of the aphid Macrosiphum euphorbiae
(Thomas, 1878), an important pest of several crops, belonging to a
genus that has been up to date scarcely studied at a cytogenetic level (
Specimens of Macrosiphum euphorbiae were collected on Bellis perennis (Linnaeus, 1753) in Modena (Italy) and maintained at 22°C with 16:8 hours light/darkness on Bellis perennis
plants. Male aphids were obtained by exposing parthenogenetic females
to short photoperiods (8:16 hours light/darkness) according to
Chromosome preparations from 150 parthenogenetic females were made by spreading embryo cells, as described by
C banding treatment was performed according to the technique of
DNA extraction from aphid embryos was performed as described in
The 28S rDNA probe was obtained by PCR amplification of a 400 bp long fragment of the 28S rDNA gene using the two primers, F (5’-AACAAACAACCGATACGTTCCG) and R (5’-CTCTGTCCGTTTACAACCGAGC), designed according to the insect 28S rRNA sequences available in GenBank. The amplification mix contained 100 ng genomic DNA, 1 mM of each primer, 200 mM dNTPs and 2 U of DyNAZyme II polymerase (Finnzymes Oy). Amplification was performed using a Hybaid thermal-cycler at an annealing temperature of 60°C for 1 min with an extension time of 1 min at 72°C.
In order to test the presence of the telomeric (TTAGG)n
repeat, a probe was obtained by PCR amplification using the two
primers F (TTAGG)5 and R (CCTAA)5 in the absence of template, as
described by
The telomeric and 28S rDNA probes were labelled using the PCR DIG labelling mix (Roche) according to the Roche protocols.
Southern blotting and fluorescent in situ hybridization (FISH) were made as described by
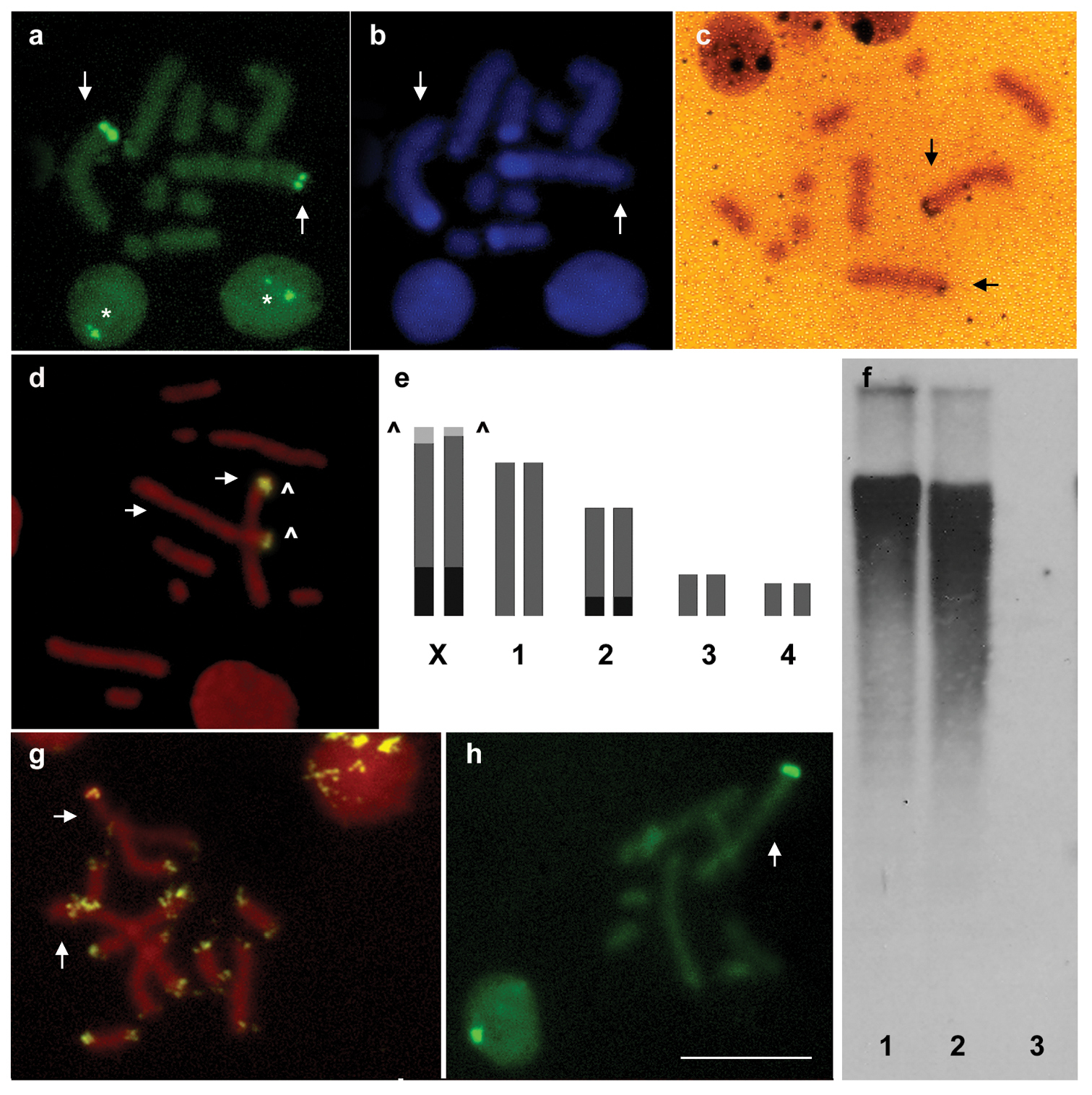
The parthenogenetic females of Macrosiphum euphorbiae showed a chromosome number of 2n=10 (Fig. 1). C banding followed by CMA3 staining showed a bright fluorescence exclusively limited to one telomere of the two longest chromosomes (Fig. 1a) that, on the basis of the comparison with male karyotype, have been identified as X chromosomes. DAPI staining showed a large heterochromatic band at the opposite end of the X chromosomes in respect to the GC-rich CMA3-stained telomere. A second heterochromatic band was observed at one telomere of the autosome pair 2 (Fig. 1b).
The overlapping between CMA3 areas and rDNA genes has been confirmed by both silver staining (Fig. 1c) and FISH with the 28S rDNA probe (Fig. 1d) showing an exclusive localization of the rDNA genes on Macrosiphum euphorbiae X chromosomes. Silver staining, FISH and CMA3 staining demonstrated the remarkable occurrence of heteromorphism between homologous NORs on X chromosomes (Figs. 1a, c, d) allowing us to distinguish the two X chromosomes. The same heteromorphism was also evident in interphase nuclei (Fig. 1a).
The presence of the (TTAGG)n repeat has been evaluated by Southern blotting and FISH. Southern blotting revealed a diffuse smear of hybridization (Fig. 1f), whereas FISH experiments with the telomeric (TTAGG)n probe showed bright FITC-fluorescent spots at the ends of all chromosomes (Fig. 1g). In the interphase nuclei of Macrosiphum euphorbiae, telomeres clustered in few highly fluorescent foci (Fig. 1g).
The male karyotype consisted of 8 autosomes and one X chromosome only (Fig. 1h). Interestingly, all the analysed males possessed the X chromosome with the larger NOR.
Macrosiphum euphorbiae chromosomes, stained with CMA3 (a) and DAPI (b) after C banding, showing heterochromatin on one telomere of the two X chromosomes and on autosome pair 2. Silver staining (c) and FISH with a 28S rDNA probe (d) evidenced heteromorphic NORs located at one telomere of each X chromosome that was also fluorescent after CMA3 staining, as summarized in the panel (e) Southern blotting (f) after digestion with XhoI of DNA samples of Acyrthosiphon pisum 1 Macrosiphum euphorbiae 2 and Drosophila melanogaster 3 together with FISH (g) assessed that the telomeric sequence (TTAGG)n constitute each chromosomal end of the Macrosiphum euphorbiae chromosomes. CMA3 staining of male chromosomes showed that all the male plates present the X chromosome with the larger NOR (h) Arrows indicate X chromosomes. Arrowheads indicate NORs. Asterisks evidence the presence of heteromorphic nucleoli in the Macrosiphum euphorbiae nuclei. Bar = 10 µm.
Currently, more than 4000 aphid species have been
described, but the chromosome number has been reported only for about
500 of them (
According to our results, Macrosiphum euphorbiae
has a chromosome number of 2n=10, which represents the typical diploid
chromosome number reported in the literature for species of this genus (
C banding carried out on Macrosiphum euphorbiae
mitotic chromosomes revealed that heterochromatin was not equilocated
on each chromosome, but limited to telomeric regions of the two X
chromosomes and to autosome pair 2. A preferential storage of
heterochromatin on X chromosomes has been previously observed in almost
all the aphid species cytogenetically studied to date with the exception
of Diuraphis noxia (Mordvilkoex Kurdjumov, 1913) (Novotna et al. 2011), but C-bands on a single autosome pair were reported only in Acyrthosiphon pisum (Harris, 1776) (
The different responses to CMA3 and DAPI staining after C banding point out a DNA composition heterogeneity of Macrosiphum euphorbiae
heterochromatin. Indeed, GCrich NOR-associated heterochromatin differs
from all other heterochromatic bands that are made by AT-rich DNAs.
This pattern of heterochromatin heterogeneity seems to be a general
characteristic of aphid chromatin since it has been described in all
species investigated so far at a cytogenetic level (
Contrary to the protocol followed for several aphid species, in order to induce a clear-cut banding pattern on Macrosiphum euphorbiae chromosomes, we modified the usual C-banding procedure making a 7 minutes long barium hydroxide treatment. Difficulties in obtaining clear-cut C-banding have been recently reported in the aphid Diuraphis noxia (Novotna et al. 2011). Interestingly, both these species belong to the tribe Macrosiphini, whereas most of the species with a clear cut banding were Aphidini. Further studies on the tribe Macrosiphini could shed light on the peculiarities of their chromatin organization, which are at the basis of this different banding propensity.
Southern blot experiments with the (TTAGG)n probe showed smears in Macrosiphum euphorbiae
genome suggesting that its telomeresare composed ofTTAGG repeats. This
result has been confirmed by FISH experiments that clearly showed a
hybridization signal on each telomere, whereas no evidence of any
interstitial labelling has been observed demonstrating that the TTAGG
repeats are restricted to the terminal regions of all aphid chromosomes.
The presence of the (TTAGG)n repeat in Macrosiphum euphorbiae further support the hypothesis that this telomeric sequence is common in aphids (
In interphase nuclei of most organisms the telomeric
regions are situated in an ordered fashion with an association to the
nuclear matrix and clustering at least in some stage of cell life (
NOR number and position have been frequently reported as
highly variable in insects, where rDNA genes have been frequently found
also on autosomes or only on autosomes as generally reported in Lepidoptera and Psylloidea (
In several aphid species silver staining revealed the
occurrence of an appreciable level of heteromorphism between homologous
NORs due to a different distribution of rDNA genes between the two X
chromosomes, but, in all the previous studies, different levels of
heteromorphism have been observed both at inter- and intra-individual
levels (
All the parthenogenetic eggs during the prophase present two X chromosomes linked by NORs (
The observation that all the Macrosiphum euphorbiae
male metaphases had an X chromosome with a large NOR evidenced a
selective bias favouring X chromosomes with larger number of rDNA genes.
This fact could be due by a non-random elimination of one X chromosome
during male determination process or by the early abortion of embryos
containing an X chromosome with few rDNA genes. Contrarily to what has
been observed in Megoura viciae (Buckton, 1876) (
The phenomenon of biased inheritance of X chromosomes in aphids seems to be controversial among aphids since observations on Sitobion fragariae
(Walker, 1848), using an X-linked polymorphic microsatellite marker,
suggested that X chromosome loss during male determination was random (
On the basis of our results on Macrosiphum euphorbiae,
we suggest that the presence of an unequal distribution of rDNA
between the two X chromosomes could affect the attachment of the X
chromosome by sticky NORs favouring the loss of the X chromosome with
few rDNA genes and a not random inheritance of the X chromosomes. On the
contrary, aphid specimens with an equal distribution of rDNA genes
could undergo a random loss of the X chromosomes during male
determination. On the basis of our hypothesis, therefore, NOR
heteromorphism does not inhibit male determination (as previously
reported by
This work is supported by the grant “F.A.R.” from the University of Modena and Reggio Emilia (M.M.).