(C) 2013 Mahua Rudra. This is an open access article distributed under the terms of the Creative Commons Attribution License 3.0 (CC-BY), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
For reference, use of the paginated PDF or printed version of this article is recommended.
Twenty five to thirty specimens each from ten populations of Mus terricolor of the Terai and the Dooars regions of the Darjeeling foothills of West Bengal were cytogenetically analyzed using C-banding. Results showed intra- and inter- population variation of C-band positive heterochromatin ranging from very large blocks to minute amounts or even complete absence of heterochromatin. Large blocks of centromeric C-bands were found in Bidhan Nagar, Garidhura, Malbazar, Nagrakata and Maynaguri populations in most of the autosomes, while the rest of the populations had large blocks of C-bands on a few autosomes only. Such intra- and inter- population variation may be due to accumulation of C-positive heterochromatin, which has not got fixed homogeneously in all autosome pairs. X-chromosomes invariably possess a C-banded short arm a telomeric C-band at the distal end of the long arm in all populations. The entire Y-chromosome was C-band positive with slight population differences in staining intensity. The results suggest quantitative as well as qualitative variation of C-positive heterochromatin.
Heterochromatin, C-banding, Mus terricolor
The earth-colored mouse Mus terricolor is a common field mouse of the Indian sub-continent infesting paddy and wheat fields and was known as Mus dunni Wroughton, 1912 until
Karyotype differentiation in Mus terricolor is due to acquisition of varying amount of constitutive heterochromatin in and around the centromere on specific autosomes. Different studies have been carried out in Mus terricolor chromosome types and their populations covering vast regions of southern, central and western part of India (
In view of the aforesaid situation, this study has been conducted to know the extent of intra- and inter- population heterochromatin variation in Mus terricolor chromosome type I from the Terai and the Dooars regions of foothills of Darjeeling in West Bengal.
The individuals of Mus terricolor were collected from paddy fields by digging burrows during harvesting season of the crop (November to December) from ten different locations of the Terai and the Dooars of foothills of Darjeeling in West Bengal, India. Three of the collection sites are in the Terai and seven collection sites are in the Dooars. The river Tista separates the Terai and the Dooars as a physical barrier. The name of the collection sites and their provisional geographical coordinates has been shown in Table 1 along with population name and number of individuals studied from each site. Animals were collected and identified by mitotic chromosome preparation. 25–30 individuals from each population were analyzed for this study. Individuals of Mus terricolor are abbreviated for convenience according to their collection localities. In the Terai region these are NXL (Naxalbari), GDH (Garidhura), BDN (Bidhan Nagar), and in the Dooars these are APD (Alipurduar), RBD (Rohimabad), KGM (Kumargram), MNG (Maynaguri), NGK (Nagrakata), MLB (Malbazar) and CBH (Cooch Behar).
Populations, collection sites, geographical coordinates and number of studied individuals of Mus terricolor.
| Populations | Collection sites | Geographical coordinates | No. of specimens |
|---|---|---|---|
| Terai region | |||
| NXL | Naxalbari | 26°41'00"N, 88°13'00"E | 30 |
| GDH | Garidhura | 26°48'24"N, 88°16'38"E | 28 |
| BDN | Bidhan Nagar | 26°16'00"N, 88°12'00"E | 28 |
| Dooars region | |||
| APD | Alipurduar | 26°31'21"N, 89°32'37"E | 25 |
| RBD | Rohimabad | 27°54'00"N, 80°30'05"E | 27 |
| KGM | Kumargram | 26°36'50"N, 89°49'30"E | 29 |
| MNG | Maynaguri | 26°33'07"N, 88°49'26"E | 25 |
| NGK | Nagrakata | 26°54'00"N, 88°50'00"E | 29 |
| MLB | Malbazar | 27°01'00"N, 89°20'17"E | 30 |
| CBH | Cooch Behar | 26°32'05"N, 89°07'12"E | 26 |
Mitotic chromosomes were prepared from bone marrow of colchicine injected mice with hypotonic treatment following air dried method after
C-banding was carried out using the BSG (Barium/Saline/Giemsa) method of
Slides were dried and incubated for 2 h at 60°C in 2 x SSC, pH 7.2 (0.3M Sodium Chloride containing 0.03 M Tri-Sodium Citrate). SSC treated slides were rinsed in distilled water and stained in 5% Giemsa, buffered with phosphate buffer (pH 6.8) for 20–30 minutes and were differentiated in distilled water, dried and mounted in DPX medium.
A minimum of 10 plates of metaphase spreads were scored for each specimen and karyotypes were prepared from selected metaphase plates. The chromosomes were numbered on the basis of euchromatic long arms as per recommendations of the Committee on Standardized Genetic Nomenclature for mice (1972).
All the individuals of Mus terricolor analyzed from ten populations of the Terai and the Dooars demonstrated the diploid number 2n=40 with all acrocentric autosomes and a large submetacentric X and a large acrocentric Y chromosomes in the complement as characteristic. No chromosomal polymorphisms like inversion and Robertsonian translocations were observed. Chromosomes prepared from each individual showed C-band staining, however, few metaphases in each slide either did not show C-band staining or has weak stain. Analyzable metaphase spreads always showed C-bands shown in the representative karyotypes from each population (Figs 1–3).
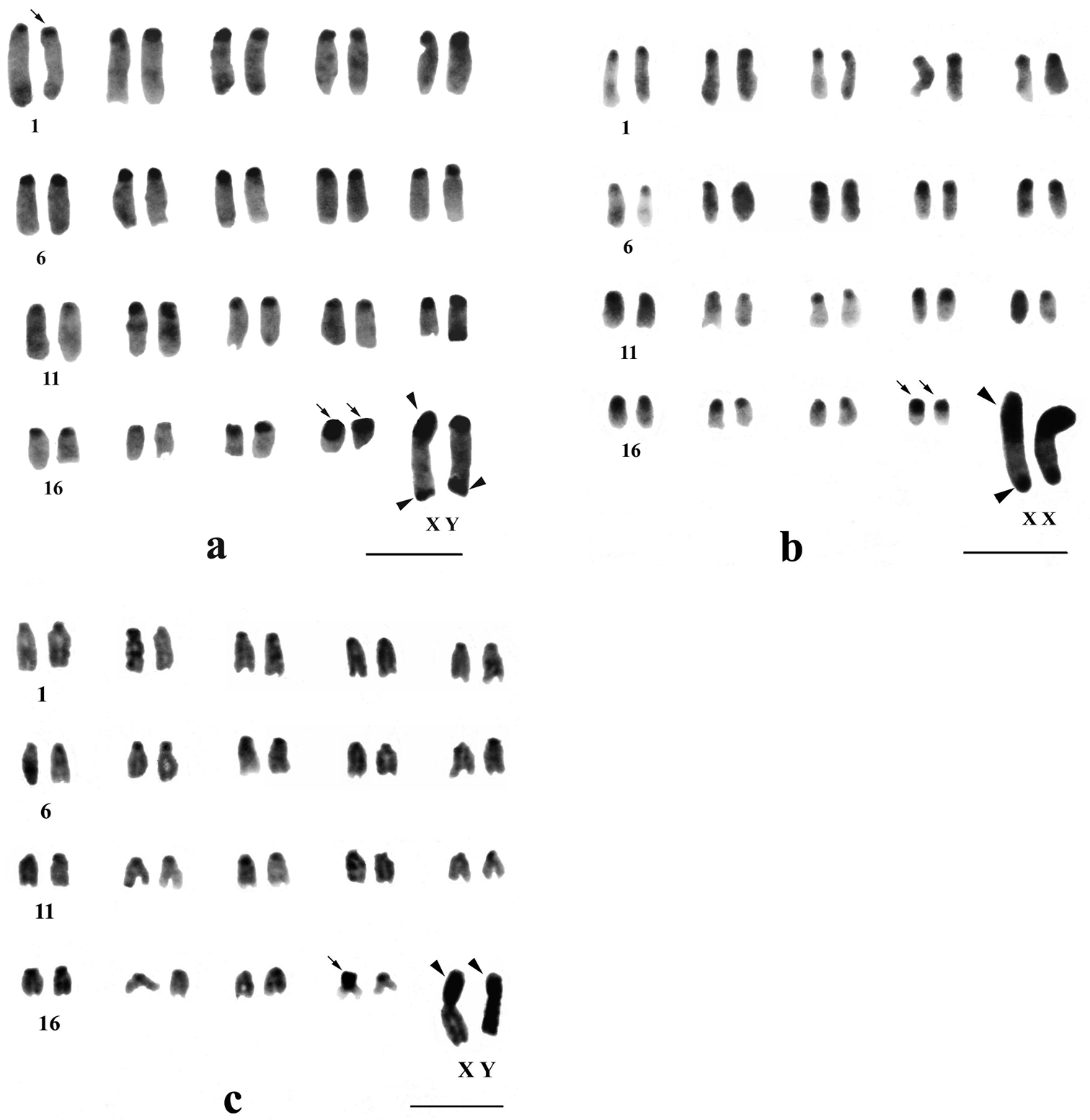
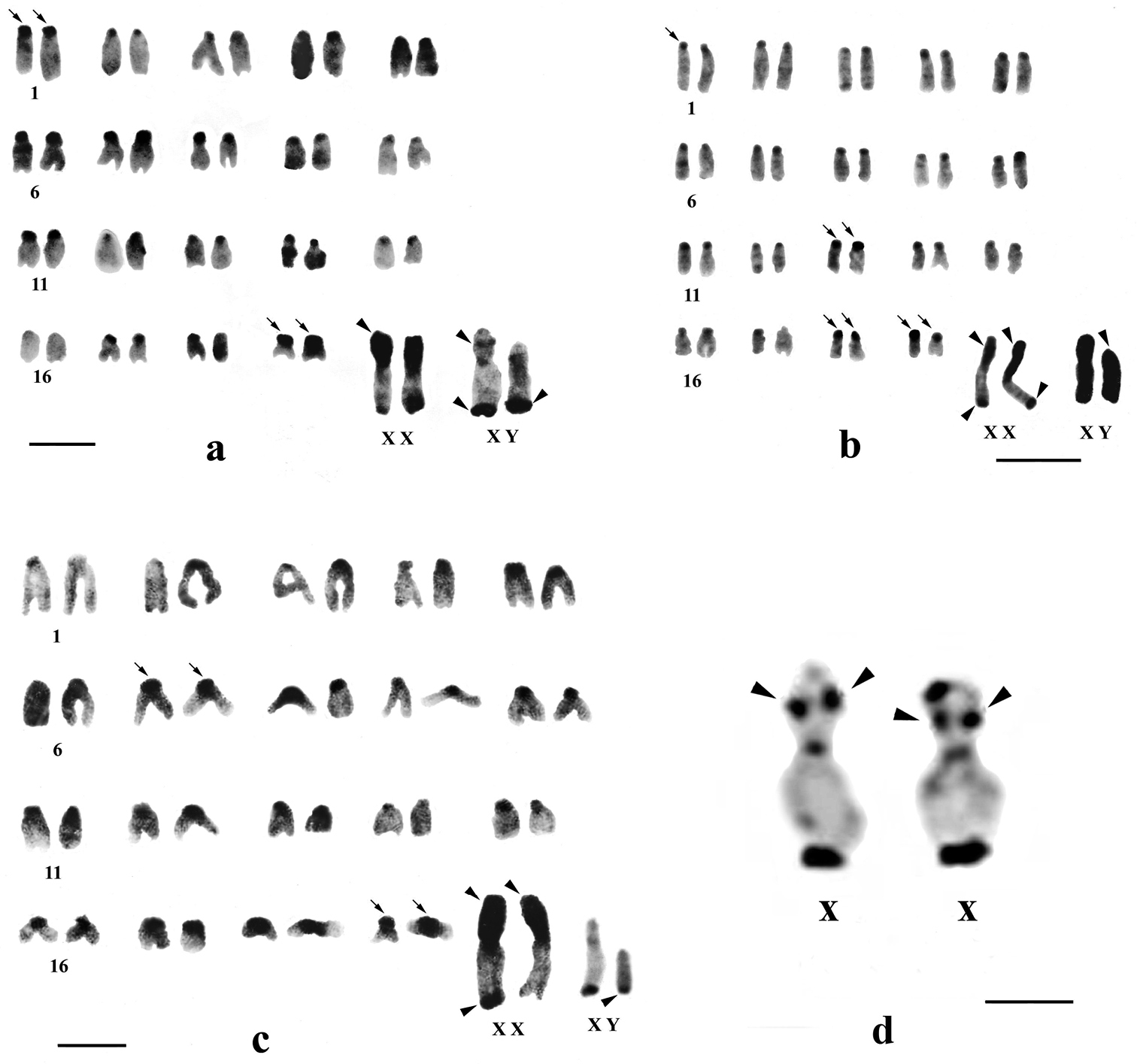
C-banded karyotypes of Mus terricolor type I from Terai populations. a NXL b BDN c GDH population d segmental C-bandon short arm of X chromosome in Mus terricolor from NXL population. Centromeric C-bands are thin arrowed, C-band in short arms of X, entire Y and telomeres of X and Y are arrow headed. Bar = 4µm.
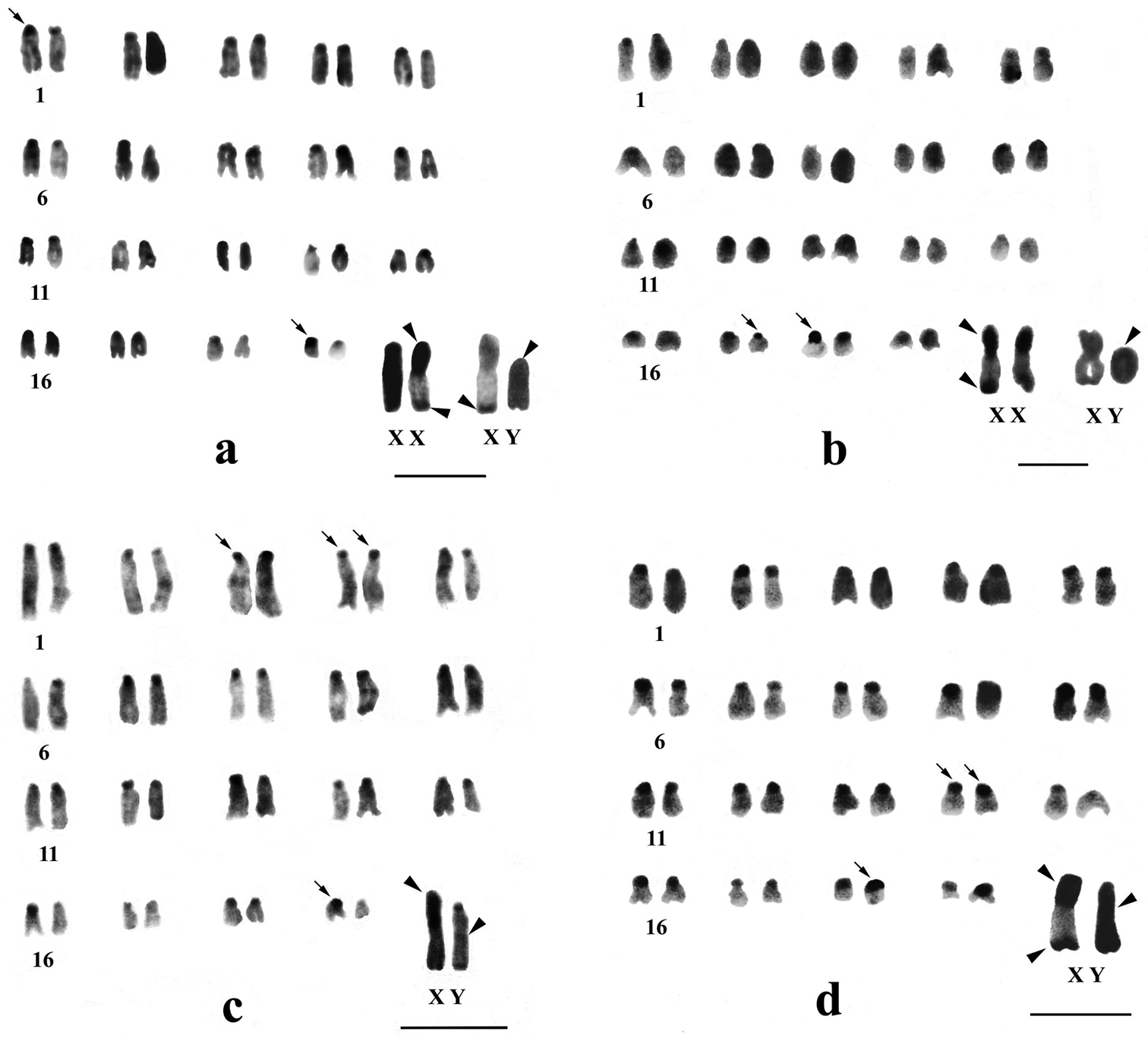
C-banded karyotypes of Mus terricolor type I from Dooars populations. a APD b RBD c KGM d MNG populations. Centromeric C-bands are thin arrowed, C-band in short arms of X, entire Y and telomeres of X and Y are arrow headed. Bar = 4µm.
C-banded karyotypes of Mus terricolor type I from Dooars populations. a MLB b NGK c CBH populations. Centromeric C-bands are thin arrowed, C-band in short arms of X, entire Y and telomeres of X and Y are arrow headed. Bar = 4µm.
Differential C-banding revealed extensive heterochromatin variation between and within populations. C-bands were found to be localized in the centromeric region of autosomes throughout the populations varying in size. According to the size of C-bands, the heterochromatin was divided into large blocks, and small to minute C-positive heterochromatin (Table 2). Results showed that individuals of BDN, GDH, MLB, NGK and MNG had large blocks of centromeric heterochromatin in most of the autosomes (Figs 1b, c, 2d, 3a, b). Moreover, the distribution of C-positive heterochromatin was not found to be homogeneous in all autosome pairs. Each chromosome of such pair was stained differentially. Autosome pair 19 consistently showed a large block of C-positive heterochromatin in almost all populations of the Terai and the Dooars with variation between the homologs of the pair (Figs 1–3). In the individuals of populations GDH, BDN, RBD, MLB, NGK and MNG large blocks of heterochromatin were also observed in chromosome 18, which was fixed in homologous condition. In contrast to other populations, NXL, RBD, APD, KGM and CBH were found to have few autosomes with prominent large blocks of C-bands (Figs 1a, 2a, b, c, 3c). Interestingly, autosome 16 was found to be C-band negative in Mus terricolor NXL of the Terai while rest of the autosomes showed moderate to prominent C-bands (Fig. 1a).
C-band variation in different populations of Mus terricolor. (s)-Heterogeneity of C-band between homologous autosome pair; SA-short arm of X; LA- long arm of X; WA- entire Y; + denotes intensity of C-band staining.
| Population | Size and location of C-positive heterochromatin in autosomes | C-positive heterochromatin in sex chromosomes | |||
| Centromere | X | Y | |||
| Large | Small to Minute | SA | LA | WA | |
| NXL | 1, 4-8, 11, 19 | 2, 3, 9, 10, 12-15, 17, 18 | +++ | + | + |
| GDH | 2(s), 3(s), 4(s), 5-8, 10-13, 15-19 | 1, 2(s), 3(s), 4(s), 9, 14 | +++ | + | ++ |
| BDN | 1-14, 17(s), 18, 19 | 15, 16, 17(s) | +++ | + | +++ |
| APD | 17(s), 19(s) | 1-16, 17(s), 18, 19(s) | +++ | + | ++ |
| RBD | 5(s), 7, 10, 11(s), 12(s), 16, 18, 19 | 1-4, 5(s), 6, 8, 9, 13-15, 17 | +++ | + | +++ |
| KGM | 17(s), 19(s) | 1-16, 17(s), 18, 19(s) | +++ | + | ++ |
| MLB | 2-10, 12-15, 18, 19 | 1, 11, 16, 17 | +++ | + | ++ |
| NGK | 2, 4, 5, 8-11, 14, 16-19 | 1, 3, 6, 7, 12, 13, 15 | +++ | + | _ |
| MNG | 1-14, 16, 18, 19 | 15, 17 | +++ | + | +++ |
| CBH | 2, 13, 16, 19 | 1, 3-12, 14, 15, 17, 18 | +++ | + | +++ |
The X and Y chromosomes of Mus terricolor were found to be consistently C-band positive in all populations, however, minute differences were observed in size and intensity of C-bands both at intra- and inter- population level (Table 2, Figs 1–3).
The short arm of X chromosomes in all populations were found to be invariably C-band positive i.e. heterochromatic while the long arms were euchromatic. The telomere of long arms revealed prominent C-band positive staining. In some individuals of NXL and BDN the C-band was found to be localized at two distinct points of short arm of X, so that the short arm was differentiated into faint and darkly stained regions with strong C-band positive distal telomere (Fig. 1a, d). One female Mus terricolor in GDH population showed interesting result. One of the two X-chromosomes in this specimen was strongly stained at the telomeric end but the other X was totally devoid of C-band positive telomeric staining, while short arm was intensly C-band positive (Fig. 1c).
The entire Y chromosome was found to be consistently C-band positive in all populations; however, some differences were noticed in the intensity of banding (Table 2). Faintly stained Y chromosome was observed in NXL, GDH, KGM and MLB populations (Figs 1a, c, 2c and 3a), while rest of the populations revealed intensely stained Y which is the characteristic of the species. Like X chromosomes, the telomeric end of the Y was also found to be C-banded with population differences.
The mouse major satellite DNA, largely present as pericentromeric constitutive heterochromatin blocks in all chromosomes except Y, is highly repetitive (
Mus terricolor is an actively speciating incipient species complex in which constitutive heterochromatin is playing a major role in karyotype differentiation (
C-band polymorphism in X chromosomes of Mus terricolor populations revealed interesting features. Two discrete heterochromatic blocks on short arms of X chromosomes in NXL and BDN (Fig. 1a, d) suggest segmental localization of heterochromatin.
The large size of the Y chromosome in Mus terricolor is due to accumulation of C-positive heterochromatin (
Populations of Mus terricolor showed prominent telomeric C-band on the long arm of X and also on acrocentric Y, but telomeric C-bands were not observed in autosomes in any population. Large prominent autosomal telomeric C-bands have been shown in the common wood mouse, Sylvaemus sylvaticus Linnaeus, 1758 by
Intra- and inter-specific karyotype evolution involving heterochromatin has been studied and discussed in many species but the evolutionary significance of heterochromatin is not established due to simultaneous involvement of chromosomal rearrangements, like inversions and Robertsonian translocations (
It can be concluded that very large to minute C-bands and even absence of C-bands in centromere of autosomes within and between populations of Mus terricolor indicates presence of differential amount of heterochromatin which might have evolved by non-reciprocal DNA turnover mechanisms in wild populations that has also been suggested by many workers (
This work has been supported by the Department of Zoology, University of North Bengal. The authors gratefully acknowledge Professor Ananda Mukhopadhyay, Department of Zoology, University of North Bengal for his valuable comments and suggestions on the language of manuscript. Authors also acknowledge Mr. Premananda Roy, Head, Chathat High School, Siliguri, District Darjeeling who has kindly read the manuscript and made valuable suggestions.