(C) 2013 Marcello Mezzasalma. This is an open access article distributed under the terms of the Creative Commons Attribution License 3.0 (CC-BY), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
For reference, use of the paginated PDF or printed version of this article is recommended.
Citation: Mezzasalma M, Guarino FM, Aprea G, Petraccioli A, Crottini A, Odierna G (2013) Karyological evidence for diversification of Italian slow worm populations (Squamata, Anguidae). Comparative Cytogenetics 7(3): 217–227. doi: 10.3897/CompCytogen.v7i3.5398
A karyological analysis on six Italian populations the slow worm (Anguis veronensis Pollini, 1818) was performed and their genetic differentiation at the mitochondrial 16S rRNA gene fragment from a Spanish sample has been assessed. The Italian populations were karyologically uniform, all showing 2n=44 elements, of which 20 were macrochromosomes and 24 microchromosomes. Comparison with literature data on Central European populations showed a difference on the morphology of the 10th chromosome pair: submetacentric in Italian populations and telocentric in the Central European ones. Our analysis showed the presence of a fragile site on chromosomes of this pair, suggesting its propensity for structural rearrangements. Analysis of the 16S rRNA gene fragment showed uniformity among Italian populations (uncorrected genetic distance of 0.4%), and their genetic distinctness from the Spanish individual (uncorrected genetic distance of 4.2%). Our results confirm the existence of two different Anguis fragilis Linnaeus, 1758 lineages, each one characterized by a different cytotype.
Karyotype, chromosome banding, 16S rRNA, Anguis, Italian Peninsula
Until recentlythere were only two recognized species of the genus Anguis Linnaeus, 1758 in the Palaeartic region: Anguis cephallonica Werner, 1894 and Anguis fragilis Linnaeus, 1758, commonly known as slow worms. The first species was considered a Mediterranean endemic restricted to the Peloponnese and some Ionian islands, while the second (Anguis fragilis) was considered a widespread taxon distributed from Western Europe to NW Iran and from the Mediterranean coast to Scandinavia, with a broad altitudinal distribution, ranging from the sea level up to 2300 m above sea level (Gasc et al. 1997,
Eleven individuals from six geographically distinct Italian localities were analysed in this work: Travacò Siccomario (Pavia, northern Italy), one male; Ancona (central Italy), two males; Valle Agricola (Caserta, southern Italy), one male and one female; Giffoni Valle Piana (Salerno, southern Italy), one male and one female; Ruvo del Monte (Potenza, southern Italy), two males and one females; Monte Cocuzzo (Cosenza, southern Italy), one males. As already successfully performed (
Total genomic DNA was extracted from blood cells using conventional phenol-chloroform method (
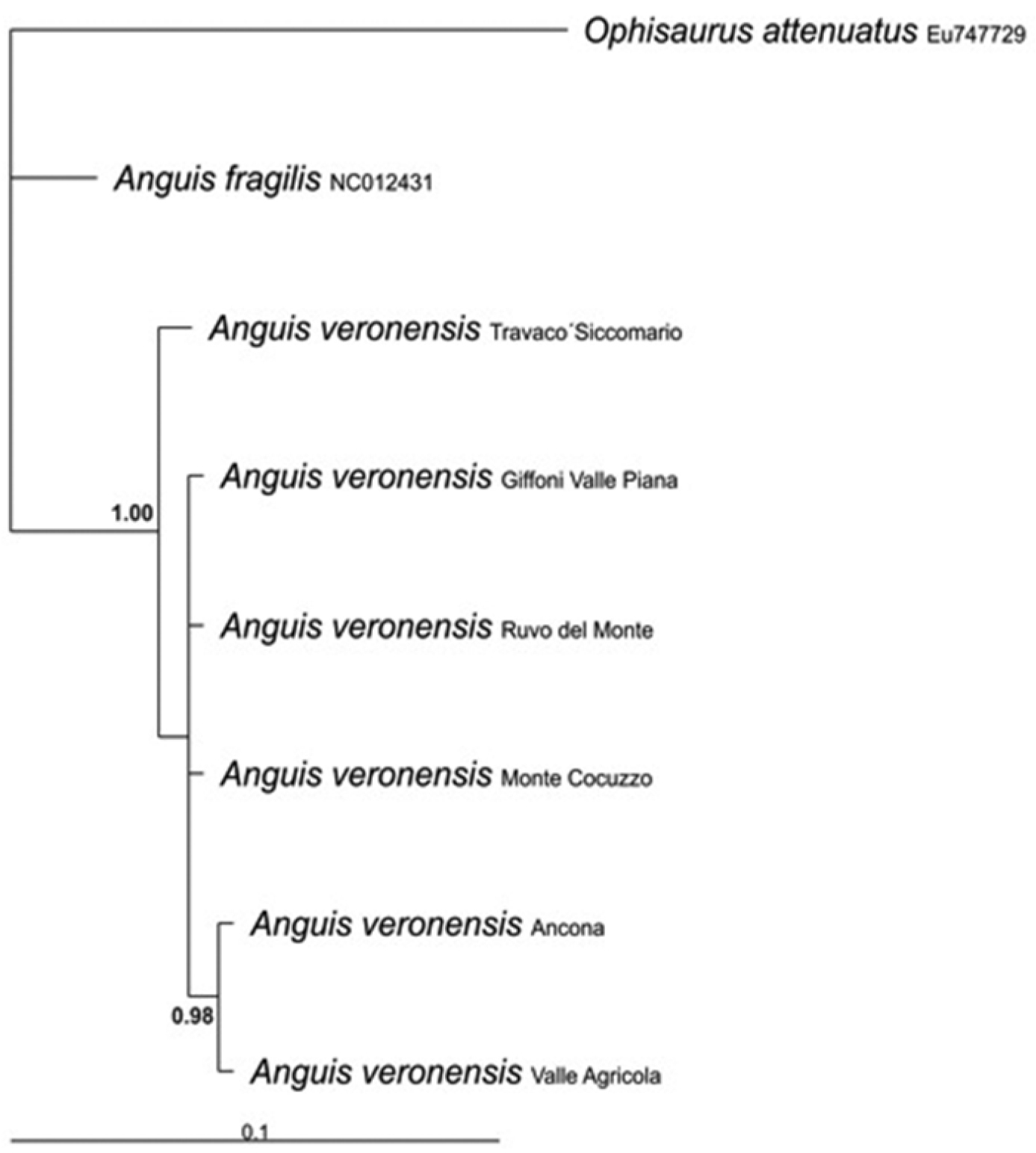
70%-majority consensus tree derived from a Bayesian inference analysis of 321 bp of the mitochondrial 16S rRNA gene. Ophisaurus attenuatus was used as outgroup. Sequences retrieved from GenBank are marked with their accession numbers.
The alignment of the analysed 16S rRNA gene fragments showed a minimum of 14 nucleotidic substitutions (11 transitions and 3 transversions) and 2 insertion/deletions between samples from Italy and Spain, corresponding to an average uncorrected genetic distance of 4.2%, thus confirming the genetic distinctiveness of the Italian populations. On the contrary, the analyzed Italian specimens were genetically very uniform and showed an intraspecific uncorrected divergence of 0.4%, whereas their uncorrected genetic distance from the outgroup, Ophisaurus attenuatus, was 12.8%. According to
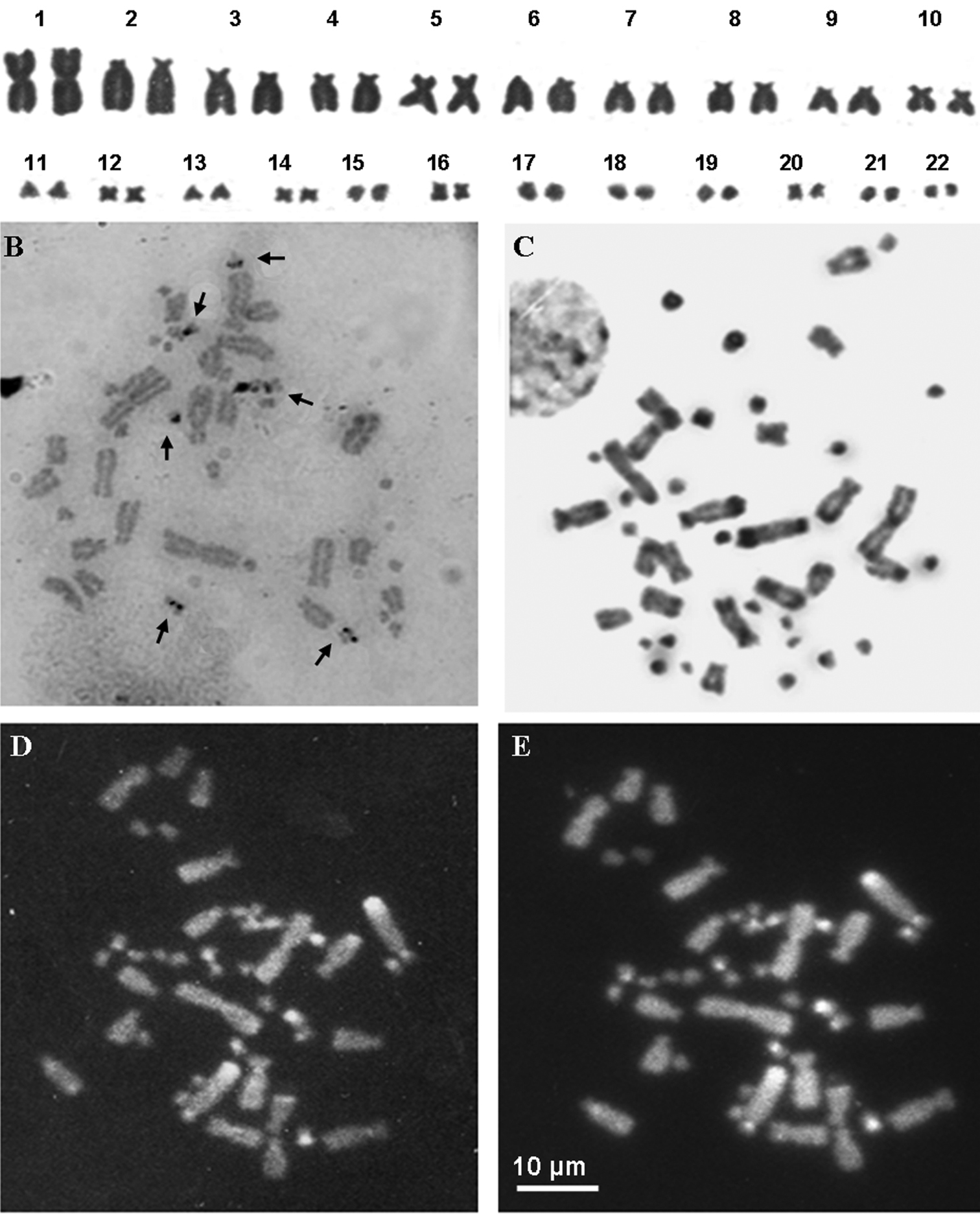
A Giemsa stained karyotype of an individual from Valle Agricola B Ag-NOR-banding stained metaphase plate of an individual from Ruvo del Monte C, D, E metaphase plate of an individual from Valle Agricola stained with C-banding (C) and C-Banding+ CMA3 (D)+ DAPI (E). Arrows in (B) point at NOR loci.
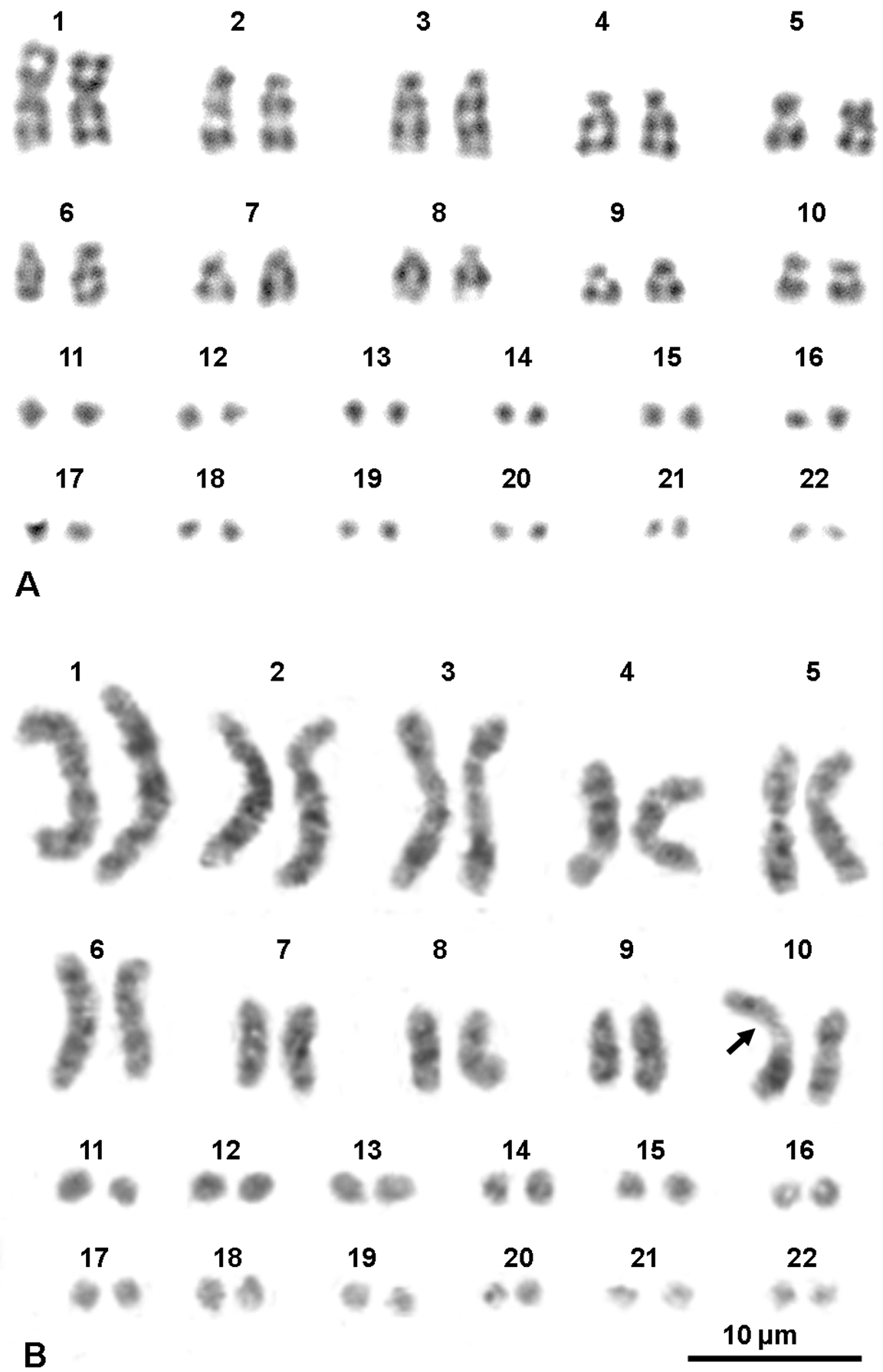
Karyotypes of a male from Travacò Siccomario (A) and of female from Ruvo del Monte (B) stained with G-banding and replication pattern, respectively. The arrow in (B) points at a fragile site.
Chromosome relative length (RL), centromeric index (CI) and chromosome shape (CS) of studied samples of Anguis veronensis. The values of RL and CI are expressed as mean ± standard deviation. Chromosome morphology was measured according to Levan et al. (1964). For the microchromosomes 11–22 only their complessive RL value is provided.
| chrom. | RL | CI | CS |
|---|---|---|---|
| 1 | 15, 2 ± 3, 5 | 0, 45 ± 3, 6 | metacentric |
| 2 | 11, 7 ± 2, 7 | 0, 10 ± 3, 8 | telocentric |
| 3 | 9, 7 ± 3, 3 | 0, 08 ± 4, 4 | telocentric |
| 4 | 7, 6 ± 3, 1 | 0, 07 ± 3, 5 | telocentric |
| 5 | 7, 4 ± 3, 4 | 0, 42 ± 3, 6 | metacentric |
| 6 | 7, 1 ± 3, 1 | 0, 07 ± 4, 1 | telocentric |
| 7 | 6, 2 ± 3, 1 | 0, 10 ± 3, 1 | telocentric |
| 8 | 5, 7 ± 2, 8 | 0, 08 ± 2, 9 | telocentric |
| 9 | 5, 1 ± 2, 7 | 0, 07 ± 4, 0 | telocentric |
| 10 | 4, 6 ± 3, 2 | 0, 34 ± 3, 8 | submetacentric |
| 11–22 | 19, 7 ± 6, 8 | – | – |
To conclude, our karyological study is consistent and provides further support for the recently discovered molecular and morphological differentiation of the Italian slow worm lineage (
Animal sampling was conducted under following authorizations: n. SCN/2D/2000/9213 of 01/06/2000 by Ministero dell’Ambiente of Italy. We are grateful to A. Gentilli, S. Scali and V. Caputo Barucchi for providing blood aliquots of the samples from Pavia and Ancona. We thank Prof. Rakesh Rastogi for his criticisms and for language revision. The work of Angelica Crottini was supported by Fundação para a Ciência e a Tecnologia SFRH/ BPD/72908/2010. We acknowledge the project “Genomics and Evolutionary Biology” cofinanced by North Portugal Regional Operational Programme 2007/2013 (ON.2 – O Novo Norte), under the National Strategic Reference Framework (NSRF), through the European Regional Development Fund (ERDF).