Citation: Walker LI, Soto MA, Spotorno ÁE (2014) Similarities and differences among the chromosomes of the wild guinea pig Cavia tschudii and the domestic guinea pig Cavia porcellus (Rodentia, Caviidae). Comparative Cytogenetics 8(2): 153–167. doi: 10.3897/CompCytogen.v8i2.7509
Cavia tschudii Fitzinger, 1867 is a wild guinea pig species living in South America that according to the analysis of mitochondrial genes is the closest wild form of the domestic guinea pig. To investigate the genetic divergence between the wild and domestic species of guinea pigs from a cytogenetic perspective, we characterized and compared the C, G and AgNOR banded karyotypes of molecularly identified Cavia tschudii and Cavia porcellus Linnaeus, 1758 specimens for the first time. Both species showed 64 chromosomes of similar morphology, although C. tschudii had four medium size submetacentric pairs that were not observed in the C. porcellus karyotype. Differences in the C bands size and the mean number of AgNOR bands between the karyotypes of the two species were detected. Most of the two species chromosomes showed total G band correspondence, suggesting that they probably represent large syntenic blocks conserved over time. Partial G band correspondence detected among the four submetacentric chromosomes present only in the C. tschudii karyotype and their subtelocentric homologues in C. porcellus may be explained by the occurrence of four pericentric inversions that probably emerged and were fixed in the C. tschudii populations under domestication. The role of the chromosomal and genomic differences in the divergence of these two Cavia species is discussed.
Karyotype, G, C and AgNOR banding, active NORs, pericentric inversions, domestication
Cavia tschudii Fitzinger, 1867 is a wild species of guinea pig (Rodentia, Caviidae) which inhabits northern Chile, southern Peru and Bolivia and northwestern Argentina (
There is a consensus that Cavia porcellus is a domestic form derived from one of the five currently recognized wild species of guinea pigs that inhabit South America (
Considering that the karyotype provides useful characters in taxonomic and systematic studies and that changes in the number and structure of chromosomes may contribute to speciation (
Skulls, skins and liver samples for DNA analysis of all the studied animals were preserved in the collection of the Laboratorio de Citogénetica de Mamíferos, Facultad de Medicina, Universidad de Chile (LCM). We examined five Cavia tschudii specimens, four males (LCM 3199b, 3110, 3080, 3225) and one female (LCM 3232), collected in the locality of Molinos, Valle de Lluta, 18°23’S, 69°45’W, Arica, I Región, Chile, and four Cavia porcellus animals, two males (LCM 2454, 3192) from the laboratory Pirbright breed, Instituto de Salud Pública, Santiago, Chile, and two females (LCM 2479, 2489) from the Andean creole breed, Arica Agromarket, Arica, Chile.
Chromosomes were obtained from marrow cells using conventional in vivo colchicine, hypotonic method, preceded by yeast injection to improve the mitotic index (
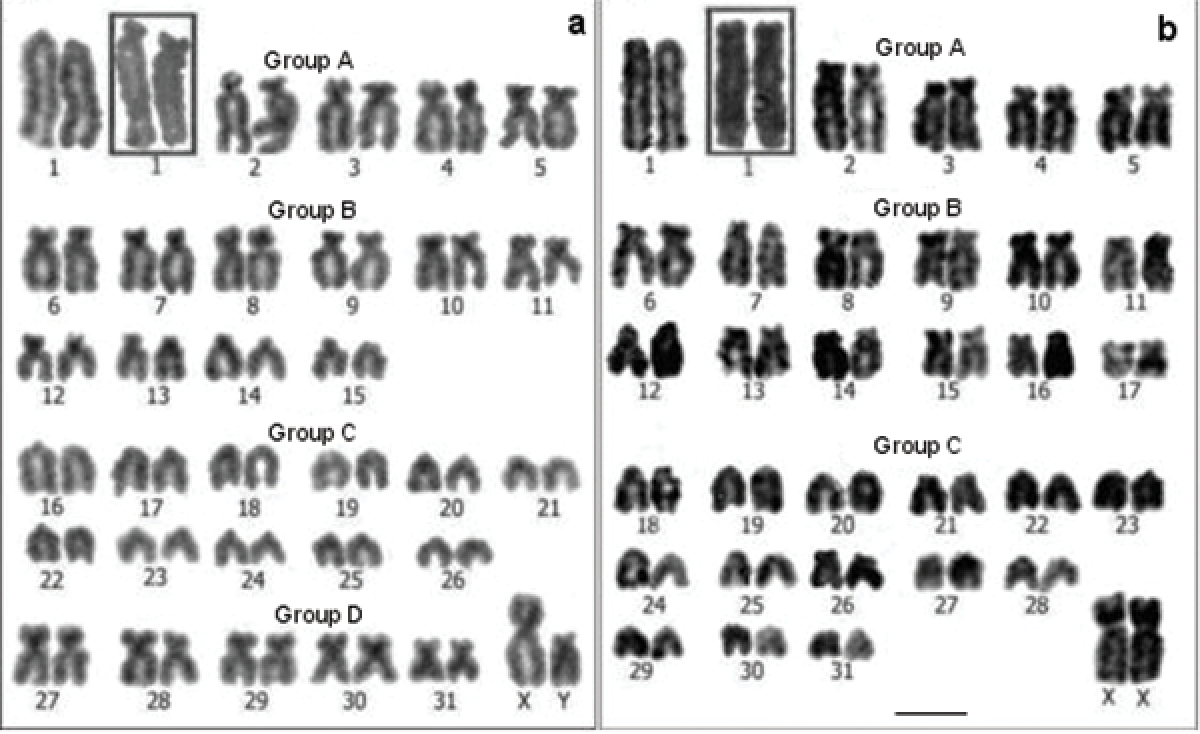
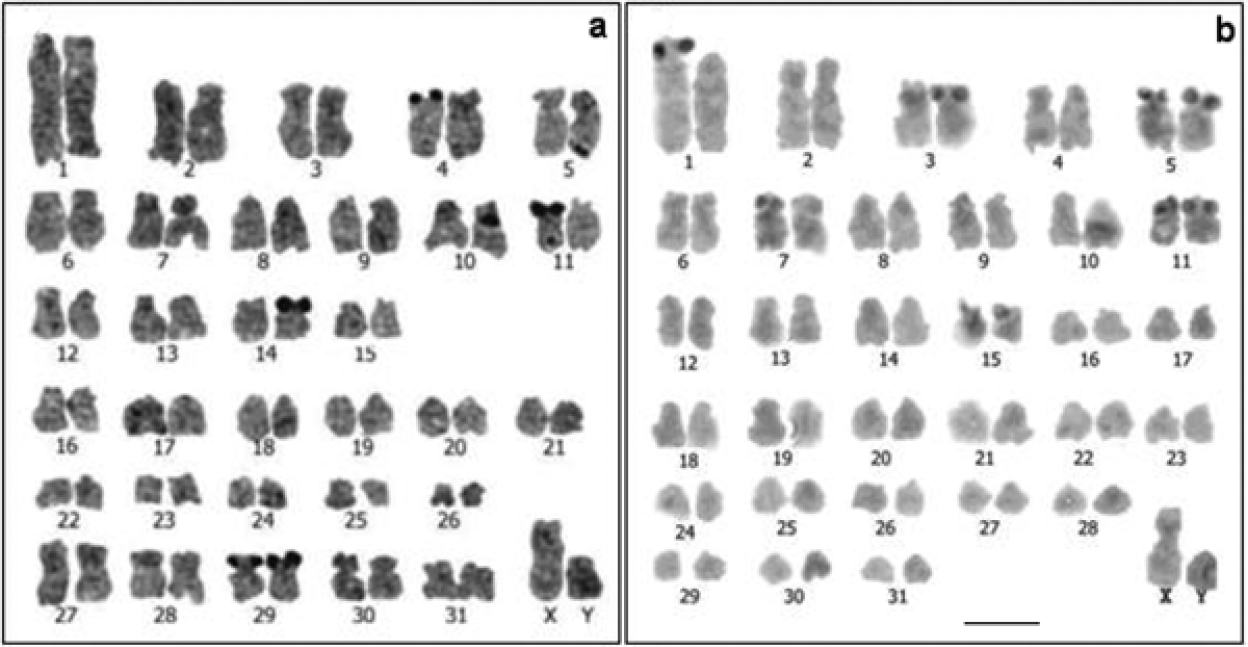
Conventional stained karyotypes: a Cavia tschudii male b Cavia porcellus female. Chromosomal pair Nº 1 shows subtelocentric morphology in other individuals of both species (a and b insets). Bar = 5 µm.
For Cavia tschudii and Cavia porcellus we consistently found a 2n = 64, FNa = 100-102; the variation in the FNa of both species was due to the polymorphism of chromosome 1 (Fig. 1). Cavia tschudii showed five pairs of submetacentric chromosomes (group D, Fig. 1a) of which four pairs (numbers 27, 28, 29 and 30) were not present in the Cavia porcellus karyotype (Fig. 1b). The X chromosome of Cavia tschudii was a large submetacentric similar to that of Cavia porcellus and the Y chromosome was a subtelocentric larger than that of Cavia porcellus (Fig. 1).
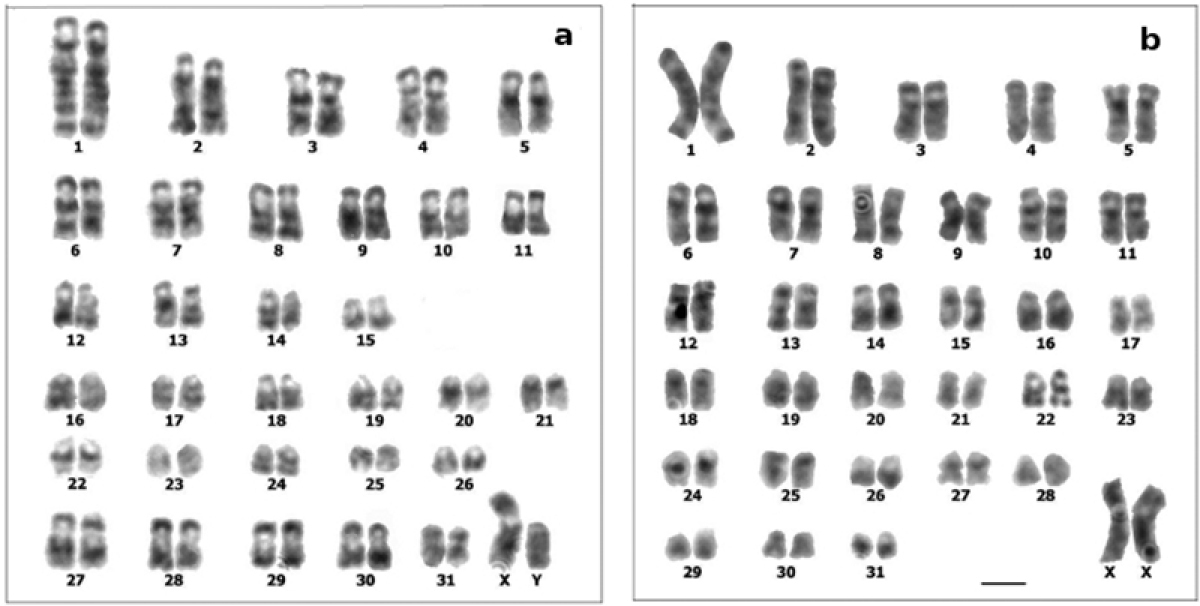
The comparison of Cavia tschudii and Cavia porcellus G-banded karyotypes revealed total correspondence for 25 of the 31 autosomal pairs and for the X chromosomes of both species (Figs 2, 3, Table 1). The four submetacentric chromosomes present only in the Cavia tschudii karyotype showed partial G band correspondence with four Cavia porcellus subtelocentric chromosomes (Fig. 6, Table 1). Only the Y chromosomes and two autosomal pairs (Cavia tschudii chromosomes 12, 13 and Cavia porcellus chromosomes 14, 18) were unique of each species karyotype (Table 1).
G-banded karyotypes: a Cavia tschudii male b Cavia porcellus female. Chromosomes numbered according to original karyotype descriptions (see Fig. 1). Bar = 5 µm.
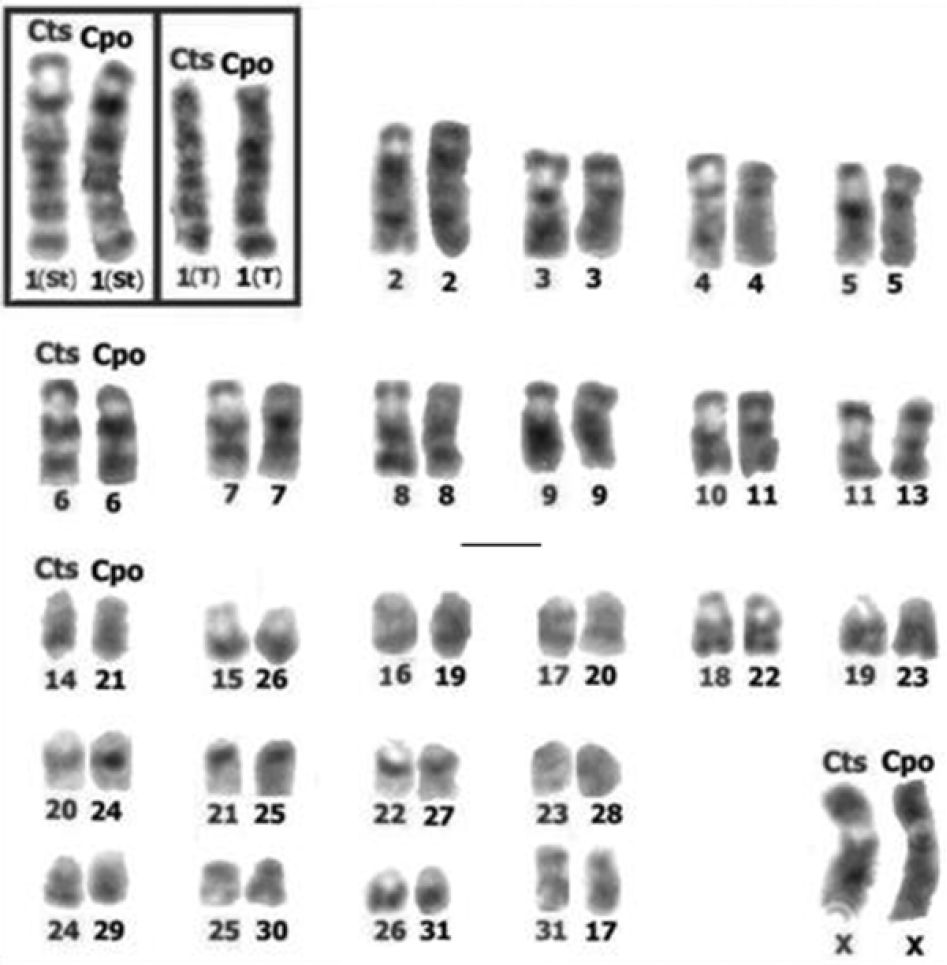
Chromosomes of Cavia tschudii (Cts) and Cavia porcellus (Cpo) with total G band correspondence. Cts chromosomes are at the left and Cpo at the right of each chromosomal group. Note that the long arms of subtelocentric (St) and telocentric (T) forms of pair 1 show total G band correspondence (inset). Chromosomes numbered according to original karyotype descriptions (see Fig. 1). Bar = 5 µm.
Correspondence of Cavia tschudii and Cavia porcellus chromosomes according to their G band patterns
| Chromosomes with total G band correspondence | ||||||||||||||||||||||||||
| Cts | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 14 | 15 | 16 | 17 | 18 | 19 | 20 | 21 | 22 | 23 | 24 | 25 | 26 | 31 | X |
| Cpo | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 11 | 13 | 21 | 26 | 19 | 20 | 22 | 23 | 24 | 25 | 27 | 28 | 29 | 30 | 31 | 17 | X |
| Chromosomes with partial G band correspondence | ||||||||||||||||||||||||||
| Cts | 27 | 28 | 29 | 30 | ||||||||||||||||||||||
| Cpo | 10 | 12 | 15 | 16 | ||||||||||||||||||||||
| Unique species chromosomes | ||||||||||||||||||||||||||
| Cts | 12 | 13 | Y | |||||||||||||||||||||||
| Cpo | 14 | 18 | Y | |||||||||||||||||||||||
1 Chromosome numbers are the one of each species karyotype (see Fig. 1); in the same column chromosomes with total or partial G band correspondence. Cts = Cavia tschudii, Cpo = Cavia porcellus.
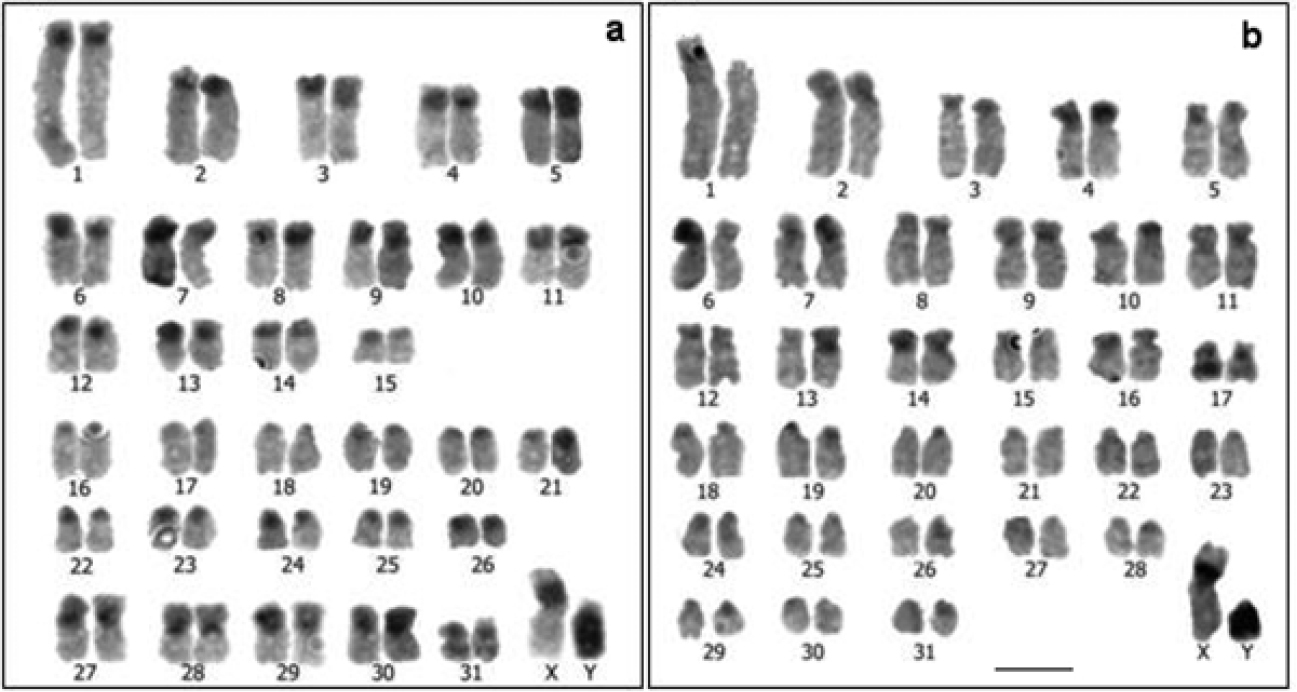
The chromosomal distribution of the C bands was similar in the karyotypes of the two species, being located preferentially in the centromeres and the short arms of the chromosomes (Fig. 4). However, the amount of constitutive heterochromatin was appreciably greater in Cavia tschudii than in Cavia porcellus, spreading over most of the short arms in several subtelocentric chromosomes (Fig. 4a). The X chromosomes of both karyotypes, equal in size, morphology and G bands (Figs 1–3), showed a C+ band in the paracentromeric region of the short arm (Fig. 4). Both Y chromosomes were completely heterochromatic, being larger the Y chromosome of Cavia tschudii than the Cavia porcellus one (Fig. 4).
C-banded karyotypes: a Cavia tschudii male b Cavia porcellus male, showing heteromorphism for chromosome 1. Most of the chromosomes of both species were tentatively identified according size and morphology. Bar = 5 µm.
Multiple AgNOR bands were detected in the karyotypes of both species, consistently located in the telomeres of several chromosomal pairs (Fig. 5). The analysis of some AgNOR banded metaphases per species indicated that the number of AgNOR bands was different between the two species and also among the individuals. Thus, the results showed that in Cavia porcellus the mean and maximum numbers of chromosomes with active NORs (5.76 and 9.0, respectively) were higher than those of Cavia tschudii (4.13 and 7.0, respectively). Moreover, when we examined all the 3.072 chromosomes from 48 metaphases of both species, each of them having 64 chromosomes, we found a total of 237 AgNOR+ sites, 138 of them located in Cavia porcellus chromosomes and 99 in Cavia tschudii chromosomes. Accordingly, the number of chromosomes bearing active NOR was significantly higher in the Cavia porcellus karyotype than in the Cavia tschudii one (χ2 = 6.956; p < 0.05; df = 1).
AgNOR-banded karyotypes: a Cavia tschudii male with four nucleolar chromosomal pairs (4, 11, 14 and 29) b Cavia porcellus male with five nucleolar chromosomal pairs (1, 3, 5, 7 and 11). The nucleolar chromosomes of both species were tentatively identified according to their size and morphology. Bar = 5 µm.
Cavia tschudii and Cavia porcellus diploid numbers (2n = 64), previously described with basic cytogenetic techniques (
The number and morphology of Cavia tschudii and Cavia porcellus chromosomes were similar to those reported for other subspecies and species of the genus Cavia. So, with the exception of Cavia intermedia Cherem, Olimpio, Ximenez, 1999, and a population of Cavia magna Ximenez, 1980, having 2n = 62 (
The analysis of the C bands showed that although they had a similar distribution in the chromosomes of the two species, they were smaller in size in the autosomes and in the Y chromosome of Cavia porcellus than in the Cavia tschudii ones, suggesting that a loss of heterochromatin occurred during the domestication process. In accordance with this result, measurements of the genome sizes of 31 hystricognath rodent species (
Five chromosomal pairs bearing NOR at the short arm telomeres were found by
The differences in the number of AgNOR bands found between the two Cavia species analyzed here and among the individuals in each of them, confirmed the tendency to variability in NOR expression usually described for mammals. It has been proposed that this variability would depend mainly on the specific metabolic demands of cells and individuals (
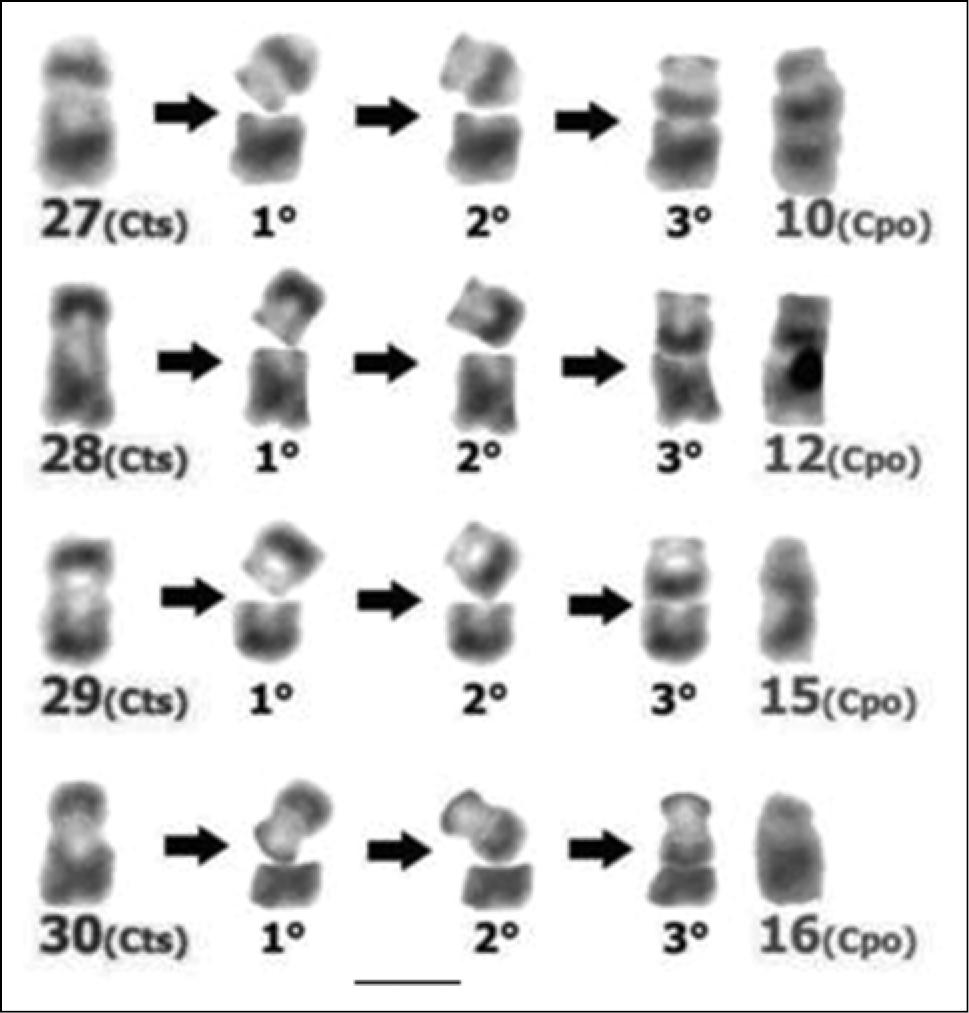
Comparison of the G-banded karyotypes of the two Cavia species included in this study revealed that most of the autosomal pairs and the X chromosomes showed total G band correspondence, suggesting that these chromosomes constitute large syntenic blocks present in the common ancestor of both species and conserved over time. The differences in morphology and the partial G band correspondences detected between four chromosomal pairs of these two species, suggest that the four submetacentric chromosomes present only in the Cavia tschudii karyotype would have suffered pericentric inversions originating the four subtelocentric chromosomes of Cavia porcellus (Fig. 6, Table 1).
Rough simulation of the changes associated with the occurrence of pericentric inversions in Cavia tschudii chromosomes. Submetacentric Cavia tschudii chromosomes (Cts, first column at the left) that originate the subtelocentric Cavia porcellus chromosomes (Cpo, last column at the right): 1°) chromosomal break, 2°) rotation of the cleaved segment, 3°) rejoining and sealing with the original segment. Chromosomes numbered according to original karyotype descriptions (see Fig. 1). Bar = 5 µm.
Cavia porcellus would be the domestic successor of Cavia tschudii from which it would have originated more than 4000 and possibly 7000 years ago (
Nevertheless, it has been demonstrated recently that the fertility of the inversion carriers is not always reduced (
A critical assessment of reproductive isolation in crosses between Cavia species as previously reported, confronts the appropriate identification of specimens, the reliability of the taxonomy at the time, and the nature of the differences eventually found. For instance, crosses between Cavia porcellus and individuals from Arequipa, Perú identified as Cavia cutleri Bennett, 1836, which correspond to the original description of Cavia cutleri based on a single specimen from Ica, Peru (see
Reproduction of wild mammal species in captivity is a difficult and not always successful task. It is even more difficult to obtain descendants from crosses between different chromosomal races or species in the laboratory (
If the analysis of the crosses realized between chromosomal races of Mus and Sorex rodents (
This study was partially financed by Project FONDECYT 1011052. We thank the Servicio Agrícola y Ganadero for the permission to capture the wild animals, Dr. Mónica Acevedo for her technical assistance and Juan Oyarce for his help in collection and maintenance of the animals.